42 Sensitivity Analysis and Robustness Checks
“Do robustness checks until you believe the result.”
— the author, having repeatedly experienced the brief joy of promising results only to watch them collapse under closer inspection, and now treating every finding as too good to be true until proven otherwise.
Sensitivity analysis and robustness checks are critical components of rigorous empirical research. They help researchers assess the stability of their findings across different model specifications, evaluate the potential impact of omitted variable bias, and provide evidence for or against causal interpretations. This chapter provides guidance on implementing various sensitivity analysis techniques, ranging from specification curve analysis to sophisticated methods for quantifying omitted variable bias.
42.1 The Philosophy of Robustness
Before diving into specific techniques, it’s worth understanding why robustness checks matter. In empirical research, we rarely have perfect certainty about the “correct” specification. Our choices about which controls to include, how to measure variables, which functional forms to use, and how to address endogeneity can all influence our results. Sensitivity analysis helps us understand whether our main conclusions depend critically on these choices, or whether they remain stable across reasonable alternatives.
A robust finding is one that persists across multiple plausible specifications. This doesn’t necessarily mean the coefficient estimate must be identical across all specifications, some variation is expected and often informative. Rather, robustness means that the key substantive conclusion (e.g., the sign, statistical significance, or economic magnitude of an effect) remains consistent despite reasonable variation in modeling choices.
42.2 Specification Curve Analysis
Specification curve analysis (also known as multiverse analysis or the specification robustness graph) provides a systematic way to examine how results vary across a large set of defensible specifications. Rather than reporting a single “preferred” specification, this approach acknowledges that multiple specifications may be equally justifiable and examines the distribution of estimates across all of them.
42.2.1 Conceptual Foundation
The specification curve approach was formalized by Simonsohn, Simmons, and Nelson (2020), though similar ideas have appeared under various names in the literature. The key insight is that researchers make many decisions when specifying a model: which controls to include, which fixed effects to add, how to cluster standard errors, etc. And these decisions can be viewed as creating a “multiverse” of possible specifications. By systematically varying these choices and examining the resulting distribution of estimates, we can assess whether our main conclusion depends on arbitrary specification choices.
A specification curve typically consists of two panels:
- The coefficient panel: Shows the point estimate and confidence interval for the main variable of interest across all specifications, typically sorted by coefficient magnitude
- The specification panel: Shows which modeling choices were made in each specification (e.g., which controls were included, which fixed effects were used)
This visualization makes it easy to see whether results are driven by particular specification choices and whether the finding is robust across the specification space.
42.2.2 The starbility Package
The starbility package provides a flexible and user-friendly implementation of specification curve analysis. It works seamlessly with various model types and allows for sophisticated customization.
42.2.2.1 Installation and Setup
# Install from GitHub
# devtools::install_github('https://github.com/AakaashRao/starbility')
library(starbility)
# Load other required packages
library(tidyverse) # For data manipulation and visualization
library(lfe) # For fixed effects models
library(broom) # For tidying model output
library(cowplot) # For combining plots42.2.2.2 Basic Specification Curve with Multiple Controls
Let’s start with an example using the diamonds dataset. This example demonstrates how to systematically vary control variables and visualize the resulting specification curve.
library(tidyverse)
library(starbility)
library(lfe)
# Load and prepare data
data("diamonds")
set.seed(43) # For reproducibility
# Create a subset for computational efficiency
# In practice, you'd use your full dataset
indices = sample(1:nrow(diamonds),
replace = FALSE,
size = round(nrow(diamonds) / 20))
diamonds = diamonds[indices, ]
# Create additional variables for demonstration
diamonds$high_clarity = diamonds$clarity %in% c('VS1','VVS2','VVS1','IF')
diamonds$log_price = log(diamonds$price)
diamonds$log_carat = log(diamonds$carat)Now let’s define our specification universe. The key is to think carefully about which specification choices are defensible and should be explored:
# Base controls: These are included in ALL specifications
# Use this for controls that you believe should always be included based on theory
base_controls = c(
'Diamond dimensions' = 'x + y + z' # Physical dimensions
)
# Permutable controls: These will be included in all possible combinations
# These are controls where theory doesn't give clear guidance on inclusion
perm_controls = c(
'Depth' = 'depth',
'Table width' = 'table'
)
# Permutable fixed effects: Different types of fixed effects to explore
# Useful when you have multiple ways to control for unobserved heterogeneity
perm_fe_controls = c(
'Cut FE' = 'cut',
'Color FE' = 'color'
)
# Non-permutable fixed effects: Alternative specifications (only one included at a time)
# Use this when you have mutually exclusive ways of controlling for something
nonperm_fe_controls = c(
'Clarity FE (granular)' = 'clarity',
'Clarity FE (binary)' = 'high_clarity'
)
# If you want to explore instrumental variables specifications
instruments = 'x + y + z'
# Add sample weights for robustness
diamonds$sample_weights = runif(n = nrow(diamonds))42.2.2.3 Custom Model Functions
One of the most powerful features of starbility is the ability to use custom model functions. This allows you to implement your preferred estimation approach, including custom standard errors, specific inference procedures, or alternative confidence intervals.
# Custom function for felm models with robust standard errors
# This function must return a vector of: c(coefficient, p-value, upper CI, lower CI)
starb_felm_custom = function(spec, data, rhs, ...) {
# Convert specification string to formula
spec = as.formula(spec)
# Estimate the model using lfe::felm
# felm is particularly useful for models with multiple fixed effects
model = lfe::felm(spec, data = data) %>%
broom::tidy() # Convert to tidy format
# Extract results for the variable of interest (rhs)
row = which(model$term == rhs)
coef = model[row, 'estimate'] %>% as.numeric()
se = model[row, 'std.error'] %>% as.numeric()
p = model[row, 'p.value'] %>% as.numeric()
# Calculate confidence intervals
# Here we use 99% CI for more conservative inference
z = qnorm(0.995) # 99% confidence level
upper_ci = coef + z * se
lower_ci = coef - z * se
# For one-tailed tests, divide p-value by 2
# Remove this if you want two-tailed p-values
p_onetailed = p / 2
return(c(coef, p_onetailed, upper_ci, lower_ci))
}
# Alternative: Custom function with heteroskedasticity-robust SEs
starb_lm_robust = function(spec, data, rhs, ...) {
spec = as.formula(spec)
# Estimate with HC3 robust standard errors
model = lm(spec, data = data)
robust_se = sandwich::vcovHC(model, type = "HC3")
# Use lmtest for robust inference
coef_test = lmtest::coeftest(model, vcov = robust_se)
row = which(rownames(coef_test) == rhs)
coef = coef_test[row, 'Estimate']
se = coef_test[row, 'Std. Error']
p = coef_test[row, 'Pr(>|t|)']
z = qnorm(0.975) # 95% confidence level
return(c(coef, p, coef + z * se, coef - z * se))
}42.2.2.4 Creating the Specification Curve
Now we can generate our specification curve with all the bells and whistles (Figure 42.1)
# Generate specification curve
# This will create plots showing how the coefficient varies across specifications
plots = stability_plot(
data = diamonds,
lhs = 'price', # Dependent variable
rhs = 'carat', # Main independent variable of interest
model = starb_felm_custom, # Use our custom model function
# Clustering and weights
cluster = 'cut', # Cluster standard errors by cut
weights = 'sample_weights', # Use sample weights
# Control variable specifications
base = base_controls, # Always included
perm = perm_controls, # All combinations
perm_fe = perm_fe_controls, # All combinations of these FEs
# Alternative: Use non-permutable FE (only one at a time)
# nonperm_fe = nonperm_fe_controls,
# fe_always = FALSE, # Set to FALSE to include specs without any FEs
# Instrumental variables (if needed)
# iv = instruments,
# Sorting and display options
sort = "asc-by-fe", # Options: "asc", "desc", "asc-by-fe", "desc-by-fe"
# Visual customization
error_geom = 'ribbon', # Display error bands as ribbons (alternatives: 'linerange', 'none')
# error_alpha = 0.2, # Transparency of error bands
# point_size = 1.5, # Size of coefficient points
# control_text_size = 10, # Size of control labels
# For datasets with fewer specifications, you might want:
# control_geom = 'circle', # Use circles instead of rectangles
# point_size = 2,
# control_spacing = 0.3,
# Customize the y-axis range if needed
# coef_ylim = c(-5000, 35000),
# Adjust spacing between panels
# trip_top = 3,
# Relative height of coefficient panel vs control panel
rel_height = 0.6
)
# Display the plots
plots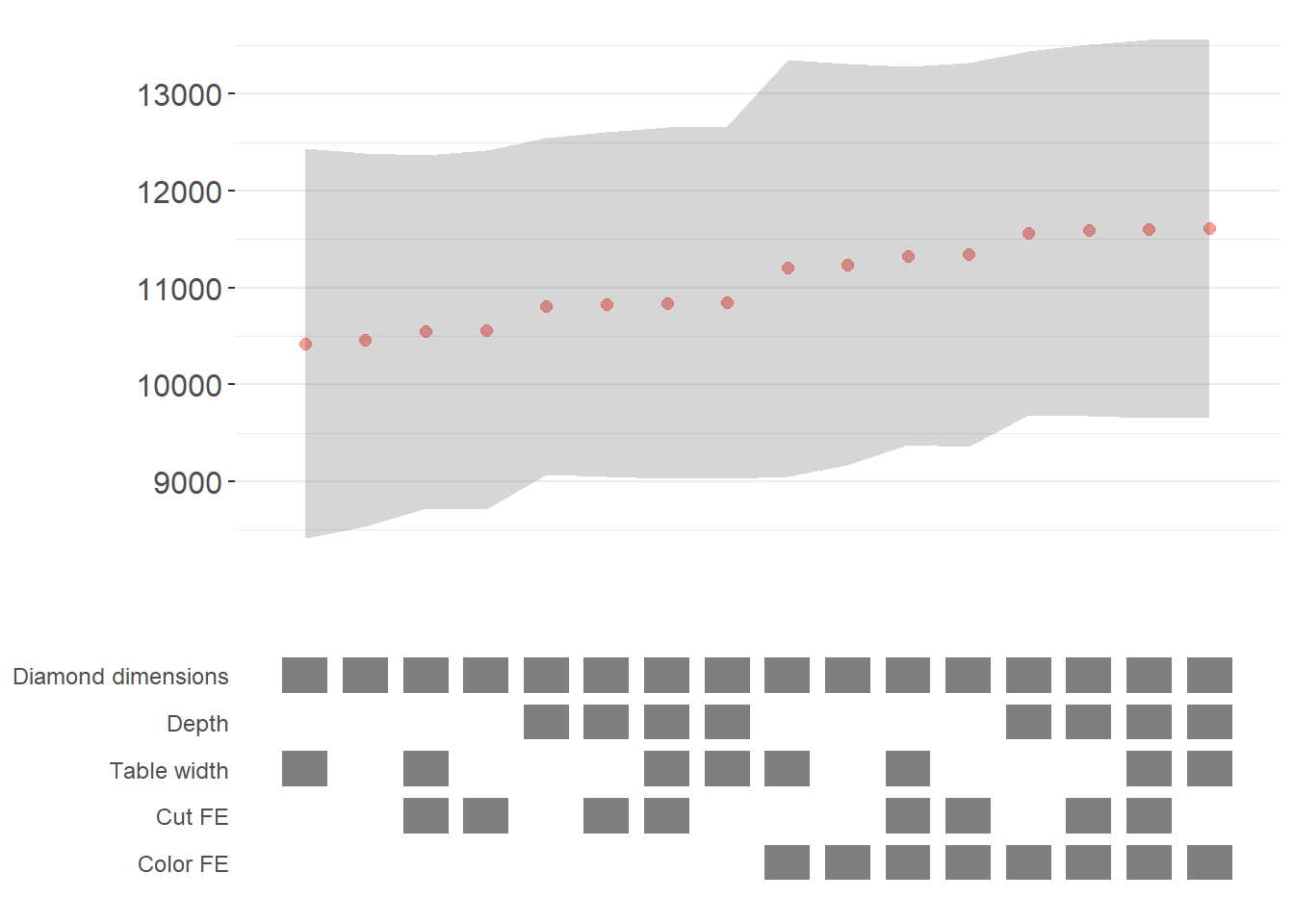
Figure 42.1: Specification curve for the effect of carat on price.
The specification curve uses color coding to indicate statistical significance:
- Red: \(p < 0.01\) (highly significant)
- Green: \(p < 0.05\) (significant)
- Blue: \(p < 0.1\) (marginally significant)
- Black: \(p > 0.1\) (not significant)
This color scheme makes it easy to see at a glance whether your finding is robust across specifications, or whether significance depends on particular specification choices.
42.2.3 Advanced Specification Curve Techniques
42.2.3.1 Step-by-Step Control of the Process
For maximum flexibility, starbility allows you to control each step of the specification curve generation process. This is useful when you want to modify the grid of specifications, use custom model functions, or create highly customized visualizations.
# Ensure high_clarity variable exists
diamonds$high_clarity = diamonds$clarity %in% c('VS1','VVS2','VVS1','IF')
# Redefine controls for this example
base_controls = c(
'Diamond dimensions' = 'x + y + z'
)
perm_controls = c(
'Depth' = 'depth',
'Table width' = 'table'
)
perm_fe_controls = c(
'Cut FE' = 'cut',
'Color FE' = 'color'
)
nonperm_fe_controls = c(
'Clarity FE (granular)' = 'clarity',
'Clarity FE (binary)' = 'high_clarity'
)
# Step 1: Create the control grid
# This generates all possible combinations of controls
grid1 = stability_plot(
data = diamonds,
lhs = 'price',
rhs = 'carat',
perm = perm_controls,
base = base_controls,
perm_fe = perm_fe_controls,
nonperm_fe = nonperm_fe_controls,
run_to = 2 # Stop after creating the grid
)
# Examine the grid structure
head(grid1, 10)
#> Diamond dimensions Depth Table width Cut FE Color FE np_fe
#> 1 1 0 0 0 0
#> 2 1 1 0 0 0
#> 3 1 0 1 0 0
#> 4 1 1 1 0 0
#> 5 1 0 0 1 0
#> 6 1 1 0 1 0
#> 7 1 0 1 1 0
#> 8 1 1 1 1 0
#> 9 1 0 0 0 1
#> 10 1 1 0 0 1
# Each row represents a different specification
# Columns indicate which controls/FEs are included (1 = yes, 0 = no)
# Step 2: Generate model expressions
# This creates the actual formula for each specification
grid2 = stability_plot(
grid = grid1, # Use the grid from step 1
data = diamonds,
lhs = 'price',
rhs = 'carat',
perm = perm_controls,
base = base_controls,
run_from = 2, # Start from step 2
run_to = 3 # Stop after generating expressions
)
# View the formulas
head(grid2, 10)
#> Diamond dimensions Depth Table width np_fe
#> 1 1 0 0 0
#> 2 1 1 0 0
#> 3 1 0 1 0
#> 4 1 1 1 0
#> 5 1 0 0 0
#> 6 1 1 0 0
#> 7 1 0 1 0
#> 8 1 1 1 0
#> 9 1 0 0 0
#> 10 1 1 0 0
#> expr
#> 1 price~carat+x+y+z|0|0|0
#> 2 price~carat+x+y+z+depth|0|0|0
#> 3 price~carat+x+y+z+table|0|0|0
#> 4 price~carat+x+y+z+depth+table|0|0|0
#> 5 price~carat+x+y+z|0|0|0
#> 6 price~carat+x+y+z+depth|0|0|0
#> 7 price~carat+x+y+z+table|0|0|0
#> 8 price~carat+x+y+z+depth+table|0|0|0
#> 9 price~carat+x+y+z|0|0|0
#> 10 price~carat+x+y+z+depth|0|0|0
# Now each row has an 'expr' column with the full model formula
# Step 3: Estimate all models
# This runs the actual regressions
grid3 = stability_plot(
grid = grid2,
data = diamonds,
lhs = 'price',
rhs = 'carat',
perm = perm_controls,
base = base_controls,
run_from = 3,
run_to = 4
)
# View estimation results
head(grid3, 10)
#> Diamond dimensions Depth Table width np_fe
#> 1 1 0 0 0
#> 2 1 1 0 0
#> 3 1 0 1 0
#> 4 1 1 1 0
#> 5 1 0 0 0
#> 6 1 1 0 0
#> 7 1 0 1 0
#> 8 1 1 1 0
#> 9 1 0 0 0
#> 10 1 1 0 0
#> expr coef p error_high error_low
#> 1 price~carat+x+y+z|0|0|0 10461.86 p<0.01 11031.84 9891.876
#> 2 price~carat+x+y+z+depth|0|0|0 10808.25 p<0.01 11388.81 10227.683
#> 3 price~carat+x+y+z+table|0|0|0 10423.42 p<0.01 10992.00 9854.849
#> 4 price~carat+x+y+z+depth+table|0|0|0 10851.31 p<0.01 11428.58 10274.037
#> 5 price~carat+x+y+z|0|0|0 10461.86 p<0.01 11031.84 9891.876
#> 6 price~carat+x+y+z+depth|0|0|0 10808.25 p<0.01 11388.81 10227.683
#> 7 price~carat+x+y+z+table|0|0|0 10423.42 p<0.01 10992.00 9854.849
#> 8 price~carat+x+y+z+depth+table|0|0|0 10851.31 p<0.01 11428.58 10274.037
#> 9 price~carat+x+y+z|0|0|0 10461.86 p<0.01 11031.84 9891.876
#> 10 price~carat+x+y+z+depth|0|0|0 10808.25 p<0.01 11388.81 10227.683
# Now includes coefficient estimates, p-values, and confidence intervals
# Step 4: Prepare data for plotting
# This creates the two dataframes needed for visualization
dfs = stability_plot(
grid = grid3,
data = diamonds,
lhs = 'price',
rhs = 'carat',
perm = perm_controls,
base = base_controls,
run_from = 4,
run_to = 5
)
coef_grid = dfs[[1]] # Data for coefficient panel
control_grid = dfs[[2]] # Data for control specification panel
head(coef_grid, 10)
#> Diamond dimensions Depth Table width np_fe
#> 1 1 0 0 0
#> 2 1 1 0 0
#> 3 1 0 1 0
#> 4 1 1 1 0
#> 5 1 0 0 0
#> 6 1 1 0 0
#> 7 1 0 1 0
#> 8 1 1 1 0
#> 9 1 0 0 0
#> 10 1 1 0 0
#> expr coef p error_high error_low
#> 1 price~carat+x+y+z|0|0|0 10461.86 p<0.01 11031.84 9891.876
#> 2 price~carat+x+y+z+depth|0|0|0 10808.25 p<0.01 11388.81 10227.683
#> 3 price~carat+x+y+z+table|0|0|0 10423.42 p<0.01 10992.00 9854.849
#> 4 price~carat+x+y+z+depth+table|0|0|0 10851.31 p<0.01 11428.58 10274.037
#> 5 price~carat+x+y+z|0|0|0 10461.86 p<0.01 11031.84 9891.876
#> 6 price~carat+x+y+z+depth|0|0|0 10808.25 p<0.01 11388.81 10227.683
#> 7 price~carat+x+y+z+table|0|0|0 10423.42 p<0.01 10992.00 9854.849
#> 8 price~carat+x+y+z+depth+table|0|0|0 10851.31 p<0.01 11428.58 10274.037
#> 9 price~carat+x+y+z|0|0|0 10461.86 p<0.01 11031.84 9891.876
#> 10 price~carat+x+y+z+depth|0|0|0 10808.25 p<0.01 11388.81 10227.683
#> model
#> 1 1
#> 2 2
#> 3 3
#> 4 4
#> 5 5
#> 6 6
#> 7 7
#> 8 8
#> 9 9
#> 10 10
# Step 5: Create the plot panels
# This generates the two ggplot objects
panels = stability_plot(
data = diamonds,
lhs = 'price',
rhs = 'carat',
coef_grid = coef_grid,
control_grid = control_grid,
run_from = 5,
run_to = 6
)
# Step 6: Combine and display
# Final step to create the complete visualization
final_plot = stability_plot(
data = diamonds,
lhs = 'price',
rhs = 'carat',
coef_panel = panels[[1]],
control_panel = panels[[2]],
run_from = 6,
run_to = 7
)42.2.3.2 Specification Curves for Non-Linear Models
Specification curve analysis is not limited to linear models. Here’s how to implement it with logistic regression (Figure 42.2).
# Create binary outcome variable
diamonds$above_med_price = as.numeric(diamonds$price > median(diamonds$price))
# Add sample weights for logit model
diamonds$weight = runif(nrow(diamonds))
# Define controls
base_controls = c('Diamond dimensions' = 'x + y + z')
perm_controls = c(
'Depth' = 'depth',
'Table width' = 'table',
'Clarity' = 'clarity'
)
lhs_var = 'above_med_price'
rhs_var = 'carat'
# Step 1: Create initial grid
grid1 = stability_plot(
data = diamonds,
lhs = lhs_var,
rhs = rhs_var,
perm = perm_controls,
base = base_controls,
fe_always = FALSE, # Include specifications without FEs
run_to = 2
)
# Step 2: Manually create formulas for logit model
# The starbility package creates expressions for lm/felm by default
# For glm, we need to create our own formula structure
base_perm = c(base_controls, perm_controls)
# Create control part of formula
grid1$expr = apply(
grid1[, 1:length(base_perm)],
1,
function(x) {
paste(
base_perm[names(base_perm)[which(x == 1)]],
collapse = '+'
)
}
)
# Complete formula with LHS and RHS variables
grid1$expr = paste(lhs_var, '~', rhs_var, '+', grid1$expr, sep = '')
head(grid1, 10)
#> Diamond dimensions Depth Table width Clarity np_fe
#> 1 1 0 0 0
#> 2 1 1 0 0
#> 3 1 0 1 0
#> 4 1 1 1 0
#> 5 1 0 0 1
#> 6 1 1 0 1
#> 7 1 0 1 1
#> 8 1 1 1 1
#> expr
#> 1 above_med_price~carat+x + y + z
#> 2 above_med_price~carat+x + y + z+depth
#> 3 above_med_price~carat+x + y + z+table
#> 4 above_med_price~carat+x + y + z+depth+table
#> 5 above_med_price~carat+x + y + z+clarity
#> 6 above_med_price~carat+x + y + z+depth+clarity
#> 7 above_med_price~carat+x + y + z+table+clarity
#> 8 above_med_price~carat+x + y + z+depth+table+clarity
# Step 3: Create custom logit estimation function
# This function estimates a logistic regression and extracts results
starb_logit = function(spec, data, rhs, ...) {
spec = as.formula(spec)
# Estimate logit model with weights, suppressing separation warning
model = suppressWarnings(
glm(
spec,
data = data,
family = 'binomial',
weights = data$weight
)
) %>%
broom::tidy()
# Extract coefficient for variable of interest
row = which(model$term == rhs)
coef = model[row, 'estimate'] %>% as.numeric()
se = model[row, 'std.error'] %>% as.numeric()
p = model[row, 'p.value'] %>% as.numeric()
# Return coefficient, p-value, and 95% CI bounds
return(c(coef, p, coef + 1.96*se, coef - 1.96*se))
}
# Generate specification curve for logit model
logit_curve = stability_plot(
grid = grid1,
data = diamonds,
lhs = lhs_var,
rhs = rhs_var,
model = starb_logit, # Use our custom logit function
perm = perm_controls,
base = base_controls,
fe_always = FALSE,
run_from = 3 # Start from estimation step
)
logit_curve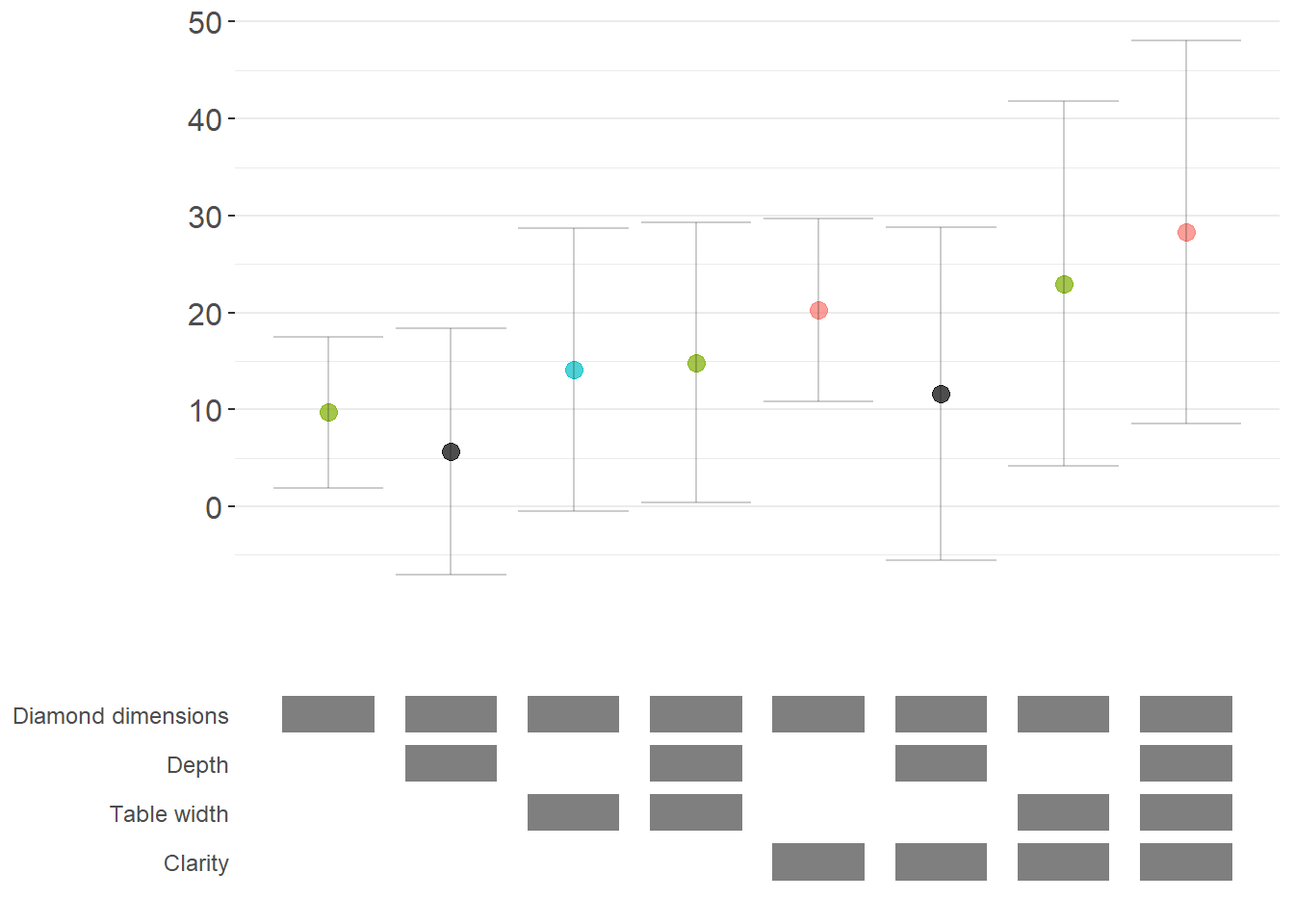
Figure 42.2: Specification curve (logit specifications).
42.2.3.3 Marginal Effects for Non-Linear Models
For non-linear models like logit or probit, we often want to report marginal effects (average marginal effects, AME) rather than raw coefficients, as they’re more interpretable. Here’s how to incorporate marginal effects into specification curves (Figure 42.3).
library(margins) # For calculating marginal effects
# Enhanced logit function with marginal effects option
starb_logit_enhanced = function(spec, data, rhs, ...) {
# Extract additional arguments
l = list(...)
get_mfx = ifelse(is.null(l$get_mfx), FALSE, TRUE) # Default to FALSE
spec = as.formula(spec)
if (get_mfx) {
# Calculate average marginal effects
model = suppressWarnings(
glm(
spec,
data = data,
family = 'binomial',
weights = data$weight
)
) %>%
margins() %>% # Calculate marginal effects
summary()
# Extract AME results
row = which(model$factor == rhs)
coef = model[row, 'AME'] %>% as.numeric() # Average Marginal Effect
se = model[row, 'SE'] %>% as.numeric()
p = model[row, 'p'] %>% as.numeric()
} else {
# Return raw coefficients (log-odds)
model = suppressWarnings(
glm(
spec,
data = data,
family = 'binomial',
weights = data$weight
)
) %>%
broom::tidy()
row = which(model$term == rhs)
coef = model[row, 'estimate'] %>% as.numeric()
se = model[row, 'std.error'] %>% as.numeric()
p = model[row, 'p.value'] %>% as.numeric()
}
# Use 99% confidence intervals for more conservative inference
z = qnorm(0.995)
return(c(coef, p, coef + z*se, coef - z*se))
}
# Generate specification curve with marginal effects
ame_curve = stability_plot(
grid = grid1,
data = diamonds,
lhs = lhs_var,
rhs = rhs_var,
model = starb_logit_enhanced,
get_mfx = TRUE, # Request marginal effects
perm = perm_controls,
base = base_controls,
fe_always = FALSE,
run_from = 3
)
ame_curve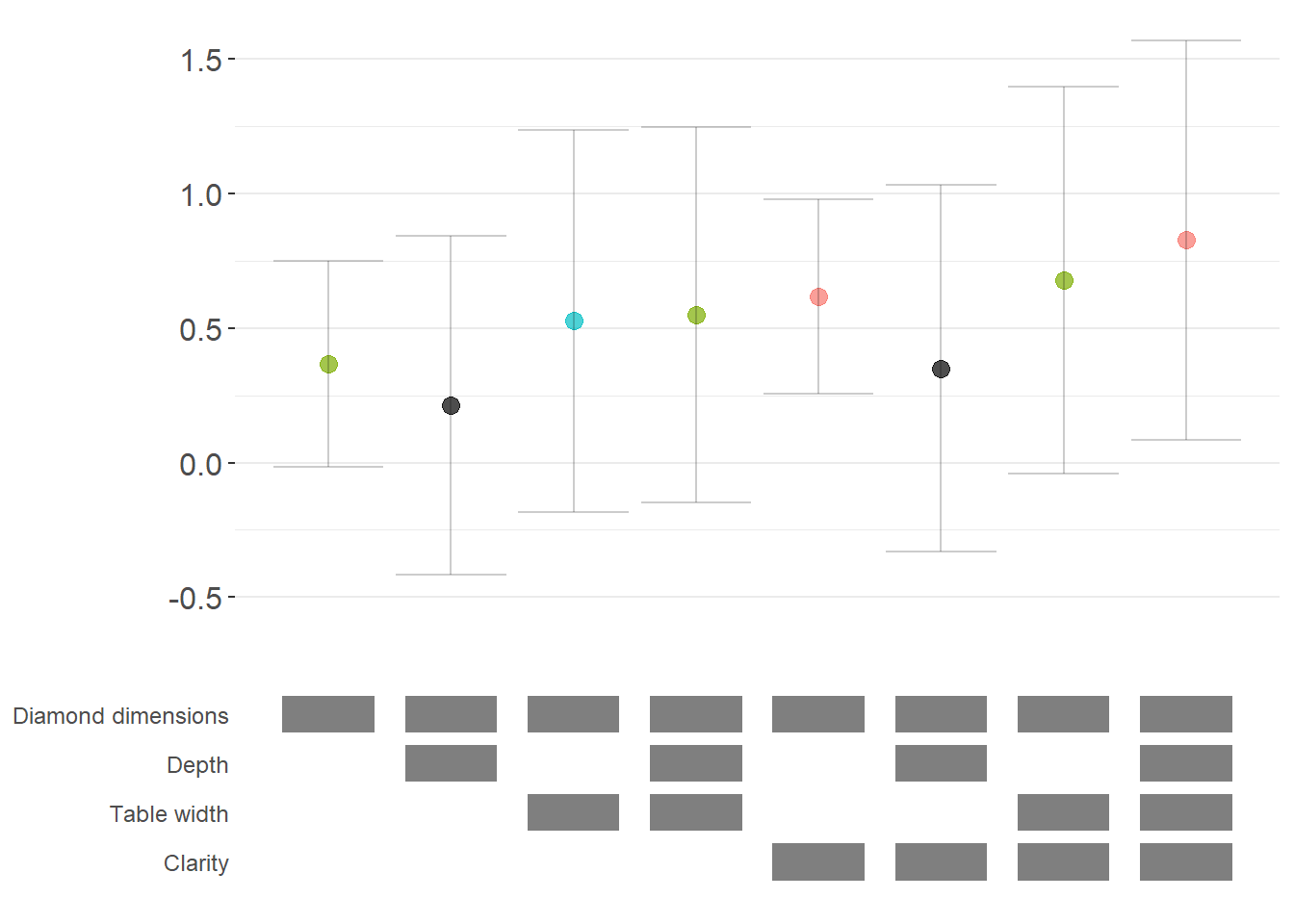
Figure 42.3: Robustness of diamond-dimension effects measured as average marginal effects.
42.2.3.4 Fully Customized Specification Curve Visualizations
When you need complete control over the appearance of your specification curve, you can extract the underlying data and create custom ggplot visualizations (Figure 42.4).
# Extract data for custom plotting
dfs = stability_plot(
grid = grid1,
data = diamonds,
lhs = lhs_var,
rhs = rhs_var,
model = starb_logit_enhanced,
get_mfx = TRUE,
perm = perm_controls,
base = base_controls,
fe_always = FALSE,
run_from = 3,
run_to = 5 # Stop before plotting
)
coef_grid_logit = dfs[[1]]
control_grid_logit = dfs[[2]]
# Define plot parameters
min_space = 0.5 # Space at edges of plot
# Create highly customized coefficient plot
coef_plot = ggplot2::ggplot(
coef_grid_logit,
aes(
x = model,
y = coef,
shape = p,
group = p
)
) +
# Add confidence interval ribbons
geom_linerange(
aes(ymin = error_low, ymax = error_high),
alpha = 0.75,
size = 0.8
) +
# Add coefficient points with color and shape by significance
geom_point(
size = 5,
aes(col = p, fill = p),
alpha = 1
) +
# Use viridis color palette (colorblind-friendly)
viridis::scale_color_viridis(
discrete = TRUE,
option = "D",
name = "P-value"
) +
# Different shapes for different significance levels
scale_shape_manual(
values = c(15, 17, 18, 19),
name = "P-value"
) +
# Reference line at zero
geom_hline(
yintercept = 0,
linetype = 'dotted',
color = 'red',
size = 0.5
) +
# Styling
theme_classic() +
theme(
axis.text.x = element_blank(),
axis.title = element_blank(),
axis.ticks.x = element_blank(),
plot.title = element_text(size = 14, face = "bold"),
plot.subtitle = element_text(size = 11)
) +
# Set axis limits
coord_cartesian(
xlim = c(1 - min_space, max(coef_grid_logit$model) + min_space),
ylim = c(-0.1, 1.6)
) +
# Remove redundant legends
guides(fill = FALSE, shape = FALSE, col = FALSE) +
# Add titles
ggtitle('Specification Curve: Effect of Carat on Above-Median Price') +
labs(subtitle = "Error bars represent 99% confidence intervals for average marginal effects")
# Create customized control specification plot
control_plot = ggplot(control_grid_logit) +
# Use diamond shapes for controls
geom_point(
aes(x = model, y = y, fill = value),
shape = 23, # Diamond shape
size = 4
) +
# Black for included, white for excluded
scale_fill_manual(values = c('#FFFFFF', '#000000')) +
guides(fill = FALSE) +
# Custom y-axis labels showing control names
scale_y_continuous(
breaks = unique(control_grid_logit$y),
labels = unique(control_grid_logit$key),
limits = c(
min(control_grid_logit$y) - 1,
max(control_grid_logit$y) + 1
)
) +
# X-axis shows specification number
scale_x_continuous(
breaks = c(1:max(control_grid_logit$model))
) +
coord_cartesian(
xlim = c(1 - min_space, max(control_grid_logit$model) + min_space)
) +
# Minimal theme for control panel
theme_classic() +
theme(
panel.grid.major.y = element_blank(),
panel.grid.minor.y = element_blank(),
axis.title = element_blank(),
axis.text.y = element_text(size = 10),
axis.ticks = element_blank(),
axis.line = element_blank()
)
# Combine plots vertically
cowplot::plot_grid(
coef_plot,
control_plot,
rel_heights = c(1, 0.5), # Coefficient plot gets more space
align = 'v',
ncol = 1,
axis = 'b' # Align bottom axes
)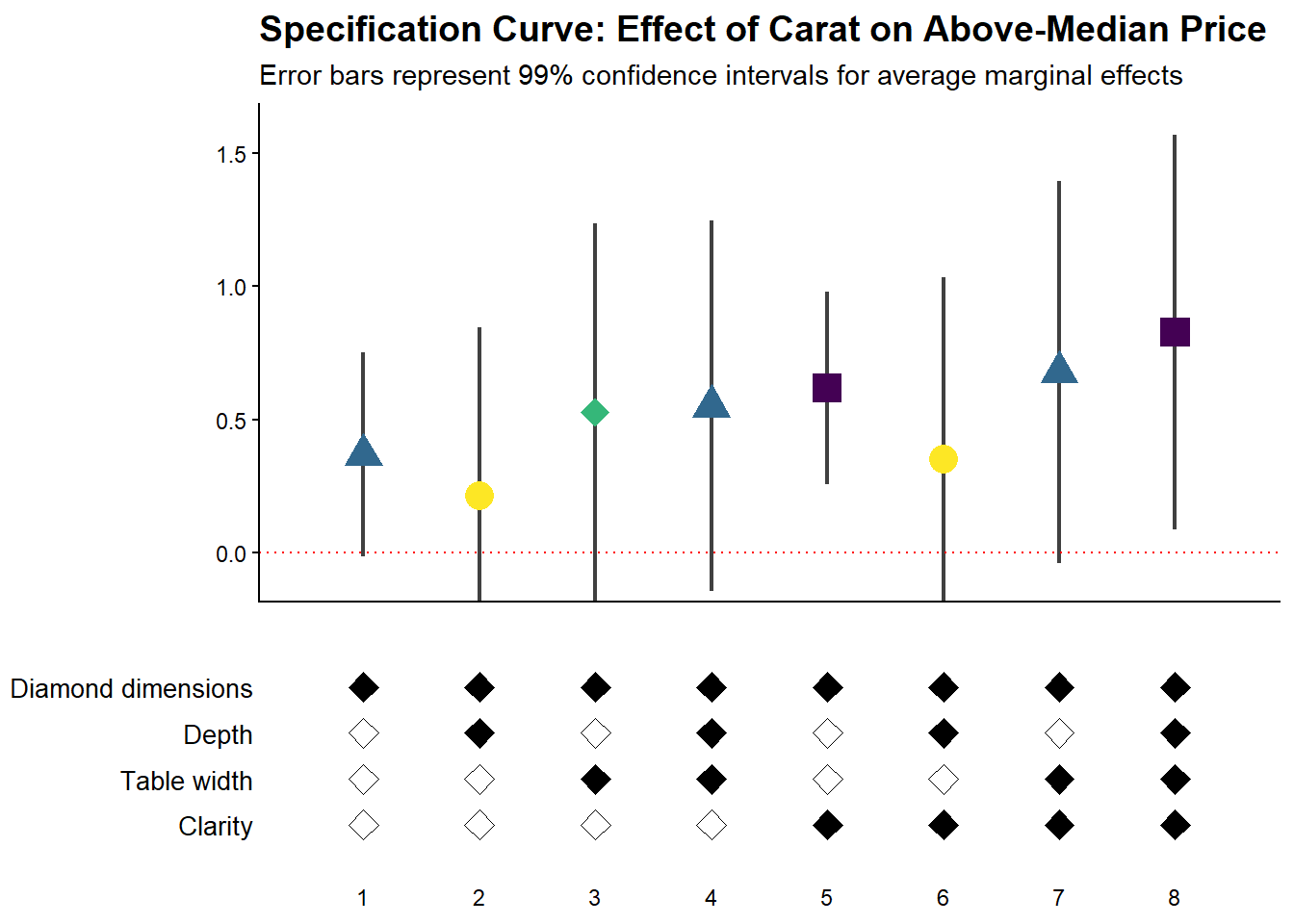
Figure 42.4: Specification curve for the effect of carat on above-median diamond price.
42.2.3.5 Comparing Multiple Model Types
A powerful extension is to compare results across different model types (e.g., logit vs. probit) in the same specification curve (Figure 42.5). This helps assess whether your findings are specific to a particular functional form assumption:
# Create custom probit estimation function
starb_probit = function(spec, data, rhs, ...) {
# Extract additional arguments
l = list(...)
get_mfx = ifelse(is.null(l$get_mfx), FALSE, TRUE)
spec = as.formula(spec)
if (get_mfx) {
# Calculate average marginal effects for probit
model = suppressWarnings(
glm(
spec,
data = data,
family = binomial(link = 'probit'), # Probit link
weights = data$weight
)
) %>%
margins() %>%
summary()
row = which(model$factor == rhs)
coef = model[row, 'AME'] %>% as.numeric()
se = model[row, 'SE'] %>% as.numeric()
p = model[row, 'p'] %>% as.numeric()
} else {
# Return raw probit coefficients
model = suppressWarnings(
glm(
spec,
data = data,
family = binomial(link = 'probit'),
weights = data$weight
)
) %>%
broom::tidy()
row = which(model$term == rhs)
coef = model[row, 'estimate'] %>% as.numeric()
se = model[row, 'std.error'] %>% as.numeric()
p = model[row, 'p.value'] %>% as.numeric()
}
# 99% confidence intervals
z = qnorm(0.995)
return(c(coef, p, coef + z * se, coef - z * se))
}
# Generate probit specification curve
probit_dfs = stability_plot(
grid = grid1,
data = diamonds,
lhs = lhs_var,
rhs = rhs_var,
model = starb_probit,
get_mfx = TRUE,
perm = perm_controls,
base = base_controls,
fe_always = FALSE,
run_from = 3,
run_to = 5
)
# Adjust model numbers for probit to plot side-by-side with logit
coef_grid_probit = probit_dfs[[1]] %>%
mutate(model = model + max(coef_grid_logit$model))
control_grid_probit = probit_dfs[[2]] %>%
mutate(model = model + max(control_grid_logit$model))
# Combine logit and probit results
coef_grid_combined = bind_rows(coef_grid_logit, coef_grid_probit)
control_grid_combined = bind_rows(control_grid_logit, control_grid_probit)
# Generate combined plots
panels = stability_plot(
coef_grid = coef_grid_combined,
control_grid = control_grid_combined,
data = diamonds,
lhs = lhs_var,
rhs = rhs_var,
perm = perm_controls,
base = base_controls,
fe_always = FALSE,
run_from = 5,
run_to = 6
)
# Add annotations to distinguish model types
coef_plot_combined = panels[[1]] +
# Vertical line separating logit and probit
geom_vline(
xintercept = max(coef_grid_logit$model) + 0.5,
linetype = 'dashed',
alpha = 0.8,
size = 1
) +
# Label for logit models
annotate(
geom = 'label',
x = max(coef_grid_logit$model) / 2,
y = 1.8,
label = 'Logit models',
size = 6,
fill = '#D3D3D3',
alpha = 0.7
) +
# Label for probit models
annotate(
geom = 'label',
x = max(coef_grid_logit$model) + max(coef_grid_probit$model) / 2,
y = 1.8,
label = 'Probit models',
size = 6,
fill = '#D3D3D3',
alpha = 0.7
) +
coord_cartesian(ylim = c(-0.5, 1.9))
control_plot_combined = panels[[2]] +
geom_vline(
xintercept = max(control_grid_logit$model) + 0.5,
linetype = 'dashed',
alpha = 0.8,
size = 1
)
# Display combined plot
cowplot::plot_grid(
coef_plot_combined,
control_plot_combined,
rel_heights = c(1, 0.5),
align = 'v',
ncol = 1,
axis = 'b'
)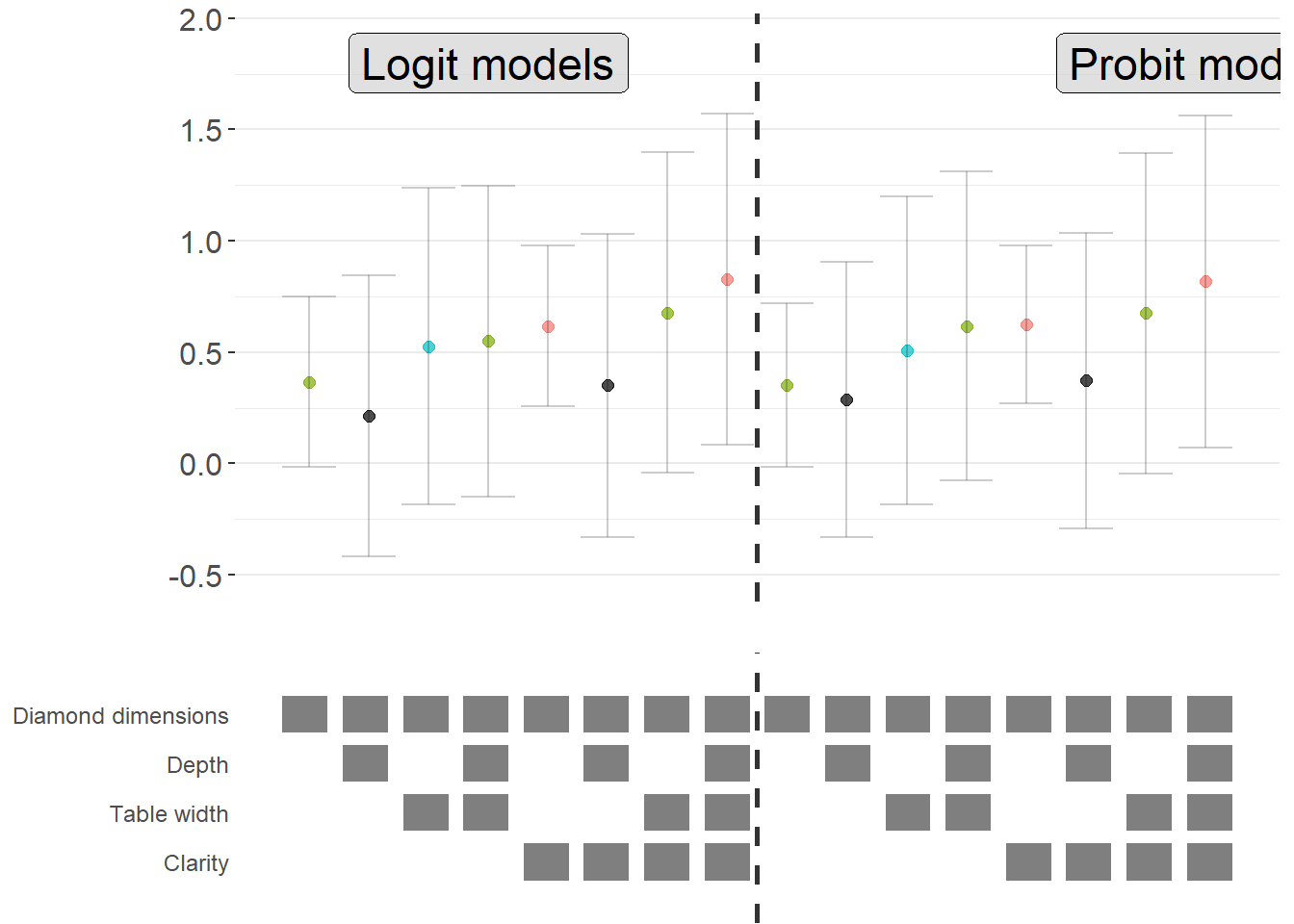
Figure 42.5: Comparison of average marginal effects across logit and probit model specifications.
42.2.4 The specr Package
The specr package provides an alternative implementation of specification curve analysis that focuses on concise specifications of the model space. Instead of defining a custom estimation function, the analyst specifies the sets of possible outcomes, focal predictors, controls, and sample restrictions. The package then enumerates all admissible specifications, estimates them, and offers summary and plotting methods.
In contrast to starbility, which is designed around user-supplied estimation functions and highly customized plotting, specr aims for a compact workflow that covers a broad range of standard models.
42.2.4.1 Preparing the Diamonds Example
To keep the exposition comparable to the starbility example, the same diamonds data are used. A subsample is created for speed, and log transformed variables are added to illustrate how specr can handle multiple outcomes and focal predictors.
library(tidyverse)
library(specr)
# Load data
data("diamonds", package = "ggplot2")
set.seed(43)
# Subsample for computational convenience
indices = sample(
x = seq_len(nrow(diamonds)),
size = round(nrow(diamonds) / 20),
replace = FALSE
)
diamonds_specr = diamonds[indices, ] |>
mutate(
high_clarity = clarity %in% c("VS1", "VVS2", "VVS1", "IF"),
log_price = log(price),
log_carat = log(carat)
)The key idea is now to describe the specification universe in terms of:
- possible outcome variables,
- possible focal predictors,
- a set of optional controls that may or may not enter the model, and
- optional sample restrictions.
specr will then traverse this design and estimate all corresponding models.
42.2.4.2 Defining and Running the Specification Curve
The central function in the package is typically called via
specr::specr(
data,
y,
x,
model,
controls,
subsets,
...
)where
-
yis a vector of outcome variable names, -
xis a vector of focal predictor names, -
modelspecifies the estimation method (for example"lm"for linear regression), -
controlsis a vector of candidate control variables, and -
subsetsis an optional list that encodes sample restrictions.
The example below sets up a relatively rich specification universe:
- outcomes: either
priceorlog_price, - focal predictor: either
caratorlog_carat, - controls: any subset of seven potential covariates.
This already yields a sizable number of specifications and illustrates how quickly the design space expands.
specs_diamonds <- specr::setup(
data = diamonds_specr,
y = c("price", "log_price"),
x = c("carat", "log_carat"),
model = "lm",
controls = c("x", "y", "z", "cut", "color")
) |>
specr::specr()
# Inspect the resulting object
summary(specs_diamonds)
#> Results of the specification curve analysis
#> -------------------
#> Technical details:
#>
#> Class: specr.object -- version: 1.0.0
#> Cores used: 1
#> Duration of fitting process: 2.73 sec elapsed
#> Number of specifications: 128
#>
#> Descriptive summary of the specification curve:
#>
#> median mad min max q25 q75
#> 1.49 171.55 -15162.63 11362.39 -0.91 6432.23
#>
#> Descriptive summary of sample sizes:
#>
#> median min max
#> 2697 2697 2697
#>
#> Head of the specification results (first 6 rows):
#>
#> # A tibble: 6 × 24
#> x y model controls subsets formula estimate std.error statistic
#> <chr> <chr> <chr> <chr> <chr> <glue> <dbl> <dbl> <dbl>
#> 1 carat price lm no covariates all price ~ … 7710. 62.3 124.
#> 2 carat price lm x all price ~ … 10340. 286. 36.2
#> 3 carat price lm y all price ~ … 9815. 282. 34.8
#> 4 carat price lm z all price ~ … 9345. 230. 40.6
#> 5 carat price lm cut all price ~ … 7816. 62.7 125.
#> 6 carat price lm color all price ~ … 8018. 63.2 127.
#> # ℹ 15 more variables: p.value <dbl>, conf.low <dbl>, conf.high <dbl>,
#> # fit_r.squared <dbl>, fit_adj.r.squared <dbl>, fit_sigma <dbl>,
#> # fit_statistic <dbl>, fit_p.value <dbl>, fit_df <dbl>, fit_logLik <dbl>,
#> # fit_AIC <dbl>, fit_BIC <dbl>, fit_deviance <dbl>, fit_df.residual <dbl>,
#> # fit_nobs <dbl>The specs_diamonds object stores one row per estimated specification, including the estimated coefficient for the focal predictor, its standard error, confidence interval, and associated \(p\) value, plus a record of which modeling choices generated that estimate.
42.2.4.3 Visualizing the Specification Curve
specr provides a plot method that produces a specification curve directly from the results object. The exact appearance depends on the package version and plotting options, but the default is typically a curve where each point corresponds to one specification, the vertical axis shows the estimated coefficient, and uncertainty intervals are plotted around each point (Figure 42.6).
# Basic specification curve plot
plot(specs_diamonds)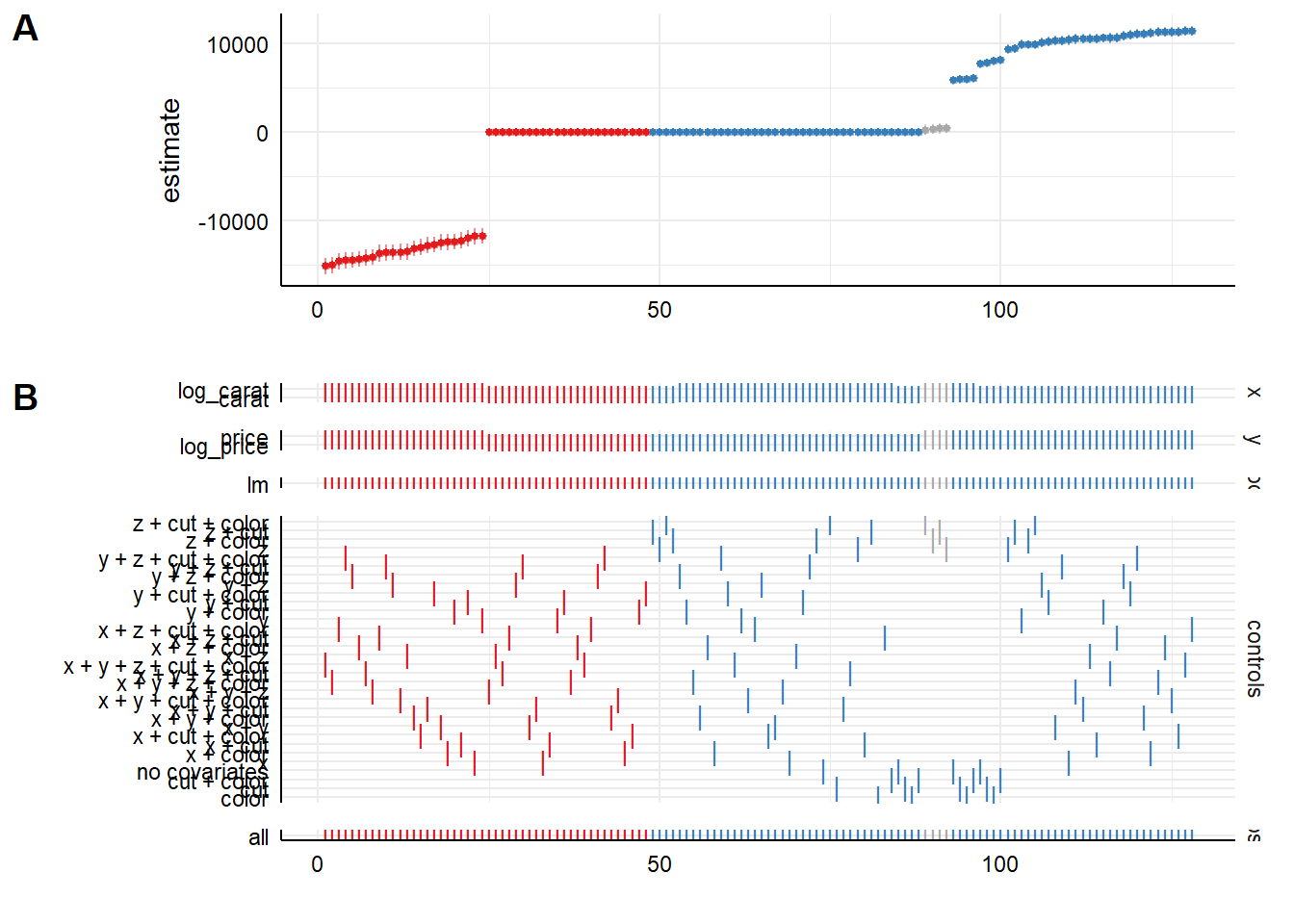
Figure 42.6: Specification-curve analysis of diamond price effects across model choices.
In this visualization (Figure 42.6), the user can typically read off:
- the range of estimates across all admissible specifications,
- how often the effect is statistically significant (for example, \(p < 0.05\)),
- whether sign changes occur as controls are added or removed, and
- how sensitive the effect size is to modeling choices.
Additional plotting options in specr generally allow the user to:
- show separate panels for different outcomes or focal predictors,
- display heatmaps of significance patterns,
- or summarize the distribution of coefficients.
The exact arguments are version dependent, so it is good practice to consult ?plot for the specr object to see the current capabilities and defaults.
42.2.4.4 Relation to the starbility Workflow
Both starbility and specr implement the same methodological idea: documenting the full set of defensible specifications and showing how the estimated effect of interest behaves across that universe.
From a practical perspective:
-
specris convenient when:- the relevant models are standard (for example linear regression, generalized linear models),
- the specification universe is naturally described in terms of outcome, focal predictor, controls, and simple subsets, and
- a concise, high level interface is preferred.
-
starbilityis preferable when:- fully custom estimation routines are needed,
- specialist estimators or complex clustering and weighting schemes are central to the analysis,
- or the user wants complete control over how coefficients, \(p\) values, and confidence intervals are computed.
In empirical work, it is entirely reasonable to begin with specr to get a quick overview of robustness patterns, then move to starbility when the analysis requires more specialized modeling choices than specr natively supports.
42.2.5 The rdfanalysis Package
While starbility is recommended for most applications, the rdfanalysis package by Joachim Gassen offers an alternative implementation with some unique features, particularly for research that follows a researcher degrees of freedom (RDF) framework.
42.2.5.1 Installation and Basic Usage
# Install from GitHub
devtools::install_github("joachim-gassen/rdfanalysis")The rdfanalysis package focuses on documenting and visualizing researcher degrees of freedom throughout the research process (Figure 42.7), from data collection to model specification:
library(rdfanalysis)
# Load example estimates from the package documentation
load(url("https://joachim-gassen.github.io/data/rdf_ests.RData"))
# Generate specification curve
# The package expects a dataframe with estimates and confidence bounds
plot_rdf_spec_curve(
ests, # Dataframe with estimates
"est", # Column name for point estimates
"lb", # Column name for lower confidence bound
"ub" # Column name for upper confidence bound
)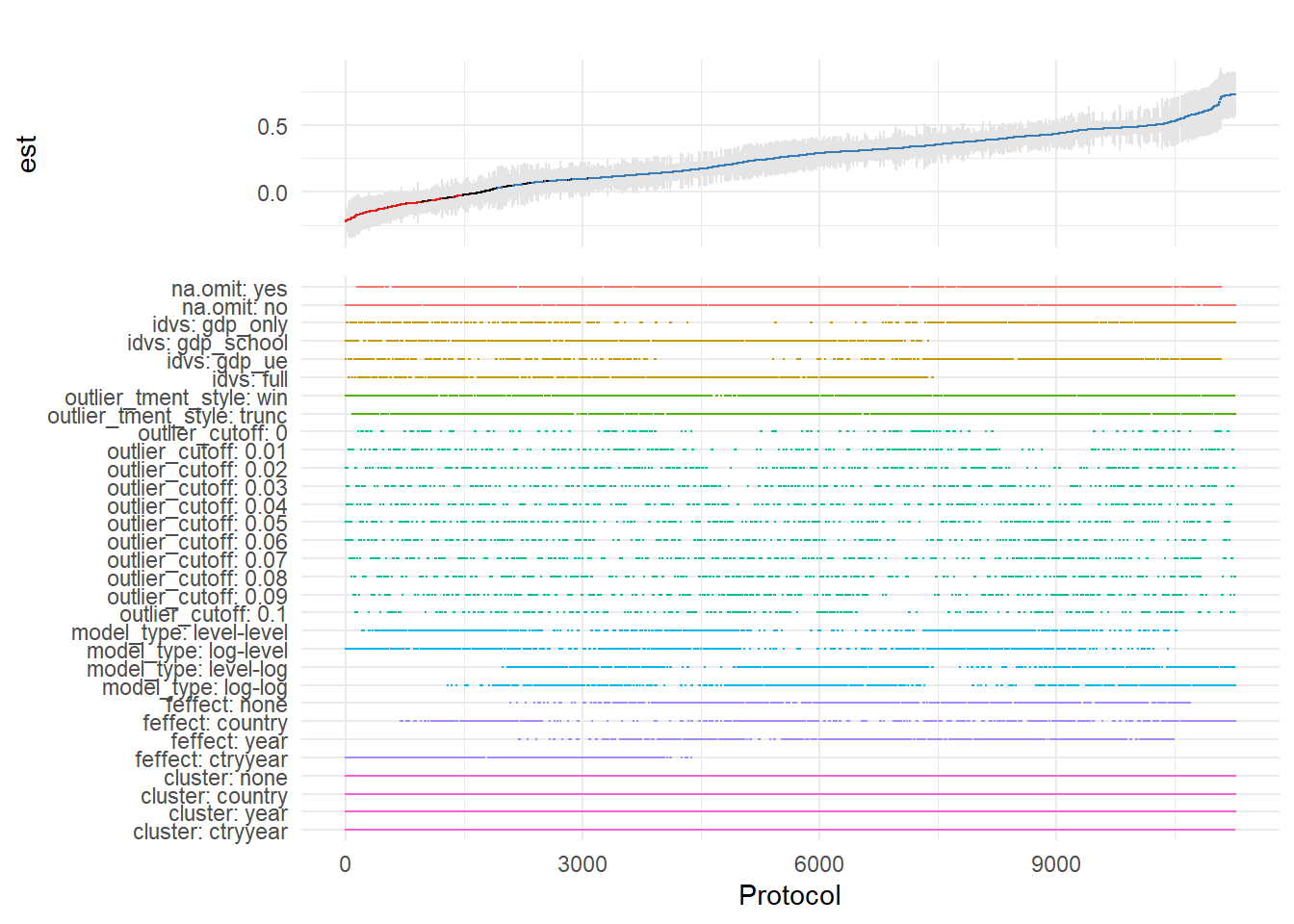
Figure 42.7: Specification curve from a multiverse analysis across analytical protocols.
This level of transparency is particularly valuable for addressing concerns about p-hacking and researcher degrees of freedom.
42.3 Coefficient Stability
Beyond specification curve analysis, another crucial aspect of sensitivity analysis is assessing whether your estimates are robust to potential omitted variable bias (Altonji, Elder, and Taber 2005). Even with comprehensive controls, unobserved confounders may threaten causal inference.
42.3.1 Theoretical Foundation: The Oster (2019) Approach
Oster’s (2019) influential paper provides a formal framework for assessing omitted variable bias. The key insight is that coefficient stability alone is insufficient, we need to consider both coefficient stability and \(R^2\) movement to evaluate the likely impact of unobservables.
The intuition is as follows:
- Coefficient stability: How much does the coefficient on your treatment variable change when you add controls?
- \(R^2\) movement: How much does the model’s explanatory power increase when you add controls?
If adding observed controls moves the \(R^2\) substantially but barely affects the coefficient, this suggests that unobservables (which might also affect \(R^2\)) are unlikely to overturn your result. Conversely, if the coefficient is very sensitive to the controls you add, unobservables might also have large effects.
Oster formalizes this by computing a parameter \(\delta\) (delta), which represents how strong the relationship between unobservables and the outcome would need to be (relative to the observables) to explain away the entire treatment effect. Higher values of \(\delta\) suggest more robust results.
The formula for calculating the bias-adjusted treatment effect is:
\[ \beta^* = \tilde{\beta} - \delta \times [\dot{\beta} - \tilde{\beta}] \times \frac{R_{max} - \tilde{R}^2}{\tilde{R}^2 - \dot{R}^2} \]
Where:
- \(\beta^*\) = bias-adjusted treatment effect
- \(\dot{\beta}\) = coefficient from short regression (without controls)
- \(\tilde{\beta}\) = coefficient from full regression (with controls)
- \(\dot{R}^2\) = \(R^2\) from short regression
- \(\tilde{R}^2\) = \(R^2\) from full regression
- \(R_{max}\) = hypothetical maximum \(R^2\) (often set to 1 or a realistic upper bound)
- \(\delta\) = proportional selection on unobservables (assumed relationship)
42.3.2 The robomit Package
The robomit package provides a straightforward implementation of Oster’s method:
library(robomit)
# Calculate bias-adjusted treatment effect using Oster's method
# This function estimates beta* under different assumptions about delta
o_beta_results = o_beta(
y = "mpg", # Dependent variable
x = "wt", # Treatment variable of interest
con = "hp + qsec", # Control variables (covariates)
delta = 1, # Proportional selection assumption
# delta = 1 means unobservables are as important as observables
# delta = 0 means no omitted variable bias
# delta > 1 means unobservables are more important than observables
R2max = 0.9, # Maximum R-squared achievable
# Common choices: 1.0 (theoretical max),
# 1.3*R2_full (30% improvement over full model),
# or domain-specific reasonable maximum
type = "lm", # Model type: "lm" for OLS, "logit" for logistic
data = mtcars # Dataset
)The function returns:
Coefficient from short model (without controls)
Coefficient from full model (with controls)
Bias-adjusted coefficient under delta and \(R^2\) max assumptions
The value of delta needed to drive the coefficient to zero
print(o_beta_results)
#> # A tibble: 10 × 2
#> Name Value
#> <chr> <dbl>
#> 1 beta* -2.00
#> 2 (beta*-beta controlled)^2 5.56
#> 3 Alternative Solution 1 -7.01
#> 4 (beta[AS1]-beta controlled)^2 7.05
#> 5 Uncontrolled Coefficient -5.34
#> 6 Controlled Coefficient -4.36
#> 7 Uncontrolled R-square 0.753
#> 8 Controlled R-square 0.835
#> 9 Max R-square 0.9
#> 10 delta 1Interpretation:
If bias-adjusted beta is still large and significant, this suggests robustness to omitted variable bias
If the delta needed to explain away the effect is large (>1), this suggests you’d need very strong unobservables to overturn the result
For a more comprehensive analysis, you can explore coefficient stability across a range of delta values:
# Create a sequence of delta values to explore
delta_values = seq(0, 2, by = 0.1)
beta_results = sapply(delta_values, function(d) {
result = suppressWarnings(
o_beta(
y = "mpg",
x = "wt",
con = "hp + qsec",
delta = d,
R2max = 0.9,
type = "lm",
data = mtcars
)
)
# Extract beta* (first row)
result$Value[result$Name == "beta*"]
})
# Create visualization
stability_df = data.frame(
delta = delta_values,
beta_adjusted = beta_results
)Interpretation guidelines:
\(|\delta| < 1\): Unobservables would need to be less related to treatment/outcome than observables to overturn the result (less robust)
\(|\delta| \approx 1\): Unobservables would need to be about as related as observables (moderate robustness)
\(|\delta| > 1\): Unobservables would need to be MORE related than observables (more robust)
\(|\delta| >> 1\): Very robust to omitted variable bias
Figure 42.8 shows bias-adjusted coefficients against \(\delta\)
# Plot
library(ggplot2)
ggplot(stability_df, aes(x = delta, y = beta_adjusted)) +
geom_line(linewidth = 1) +
geom_hline(yintercept = 0, linetype = "dashed", color = "red") +
labs(
x = "Delta (strength of confounding)",
y = "Bias-adjusted coefficient (beta*)",
title = "Coefficient stability under omitted variable bias"
) +
theme_minimal()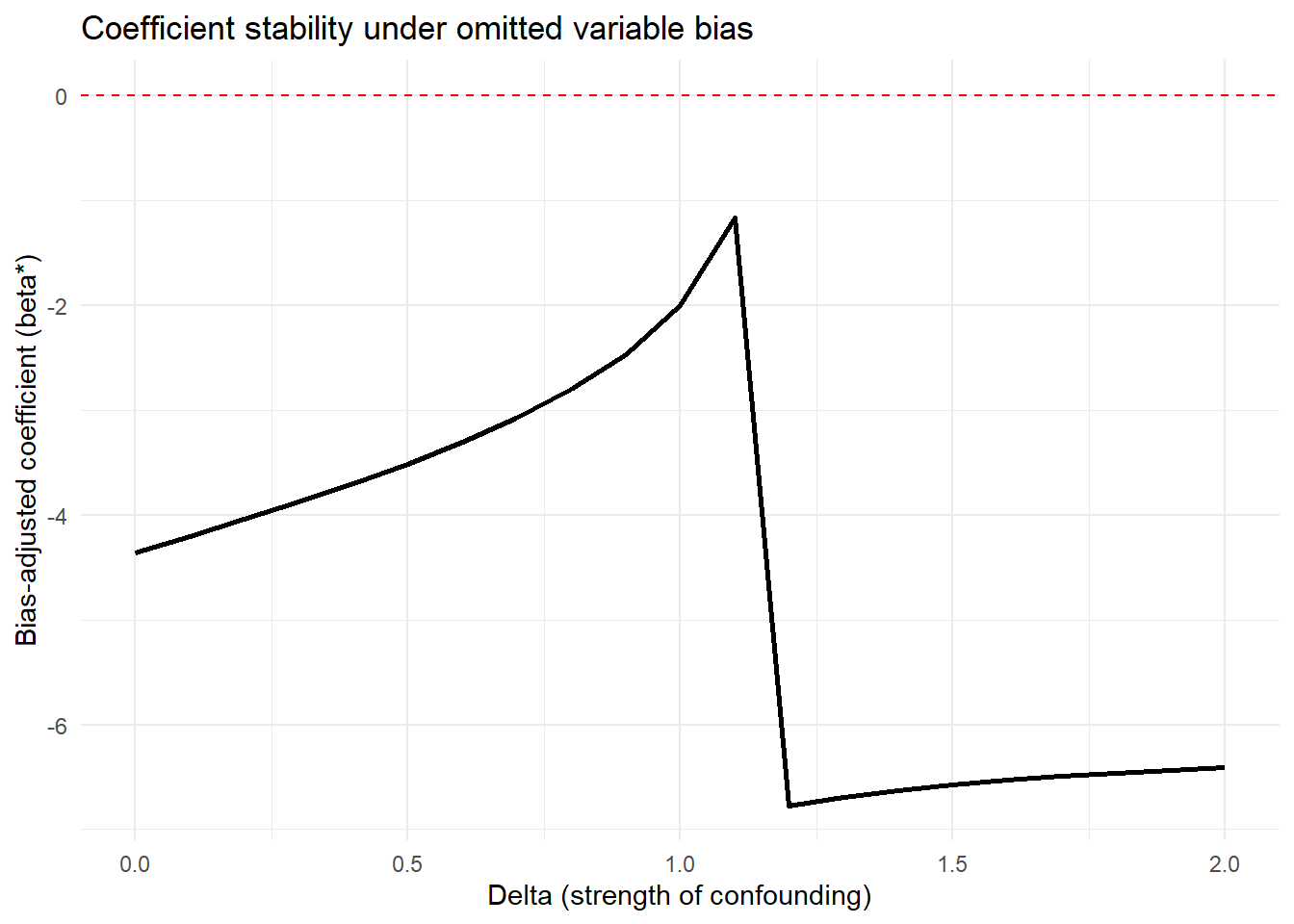
Figure 42.8: Coefficient stability under omitted-variable bias.
For more sophisticated applications with multiple treatments or different model types:
# Example with multiple treatment variables
# Useful when you have several key independent variables of interest
results_multi = lapply(c("wt", "hp"), function(treat) {
o_beta(
y = "mpg",
x = treat,
con = "qsec + gear + carb", # More extensive controls
delta = 1,
R2max = 1.0, # Theoretical maximum
type = "lm",
data = mtcars
)
})
names(results_multi) = c("wt", "hp")
print(results_multi)
#> $wt
#> # A tibble: 10 × 2
#> Name Value
#> <chr> <dbl>
#> 1 beta* 4.56
#> 2 (beta*-beta controlled)^2 68.3
#> 3 Alternative Solution 1 -6.29
#> 4 (beta[AS1]-beta controlled)^2 6.70
#> 5 Uncontrolled Coefficient -5.34
#> 6 Controlled Coefficient -3.70
#> 7 Uncontrolled R-square 0.753
#> 8 Controlled R-square 0.845
#> 9 Max R-square 1
#> 10 delta 1
#>
#> $hp
#> # A tibble: 10 × 2
#> Name Value
#> <chr> <dbl>
#> 1 beta* 0.118
#> 2 (beta*-beta controlled)^2 0.0250
#> 3 Alternative Solution 1 -0.0848
#> 4 (beta[AS1]-beta controlled)^2 0.00195
#> 5 Uncontrolled Coefficient -0.0682
#> 6 Controlled Coefficient -0.0406
#> 7 Uncontrolled R-square 0.602
#> 8 Controlled R-square 0.794
#> 9 Max R-square 1
#> 10 delta 1Comparing robustness across treatment variables helps identify which relationships are most stable.
42.3.3 The mplot Package for Graphical Model Stability
The mplot package provides complementary tools for visualizing model stability and variable selection:
# Install if needed
# install.packages("mplot")
library(mplot)
# Visualize variable importance across models
mplot::vis(
lm(mpg ~ wt + hp + qsec + gear + carb, data = mtcars),
B = 100 # Number of bootstrap samples
)
#> name prob logLikelihood
#> mpg~1 1.00 -102.38
#> mpg~wt 0.97 -80.01
#> mpg~wt+hp 0.45 -74.33
#> mpg~wt+qsec 0.36 -74.36
#> mpg~wt+qsec+gear+carb+RV 0.39 -71.46This creates plots showing:
- Which variables are selected across bootstrap samples
- Coefficient stability for each variable
- Model fit across different variable combinations
Particularly useful for:
Assessing variable importance
Understanding multicollinearity effects
Evaluating model selection stability
42.4 Quantifying Omitted Variable Bias
While Oster’s approach focuses on bias-adjusted coefficients, the Konfound framework (Narvaiz et al. 2024) takes a complementary approach by asking: “How much bias would be needed to invalidate our inference?”
42.4.1 The konfound Package
The konfound package implements sensitivity analysis for causal inferences by calculating how much unmeasured confounding would be needed to change your substantive conclusion.
library(konfound)
# Basic konfound analysis
# This calculates the amount of bias needed to invalidate your inference
pkonfound(
est_eff = 5, # Your estimated effect
std_err = 2, # Standard error of the estimate
n_obs = 1000, # Number of observations
n_covariates = 5, # Number of covariates in your model
alpha = 0.05, # Significance level
tails = 2 # Two-tailed test
)
#> Robustness of Inference to Replacement (RIR):
#> RIR = 215
#>
#> To nullify the inference of an effect using the threshold of 3.925 for
#> statistical significance (with null hypothesis = 0 and alpha = 0.05), 21.506%
#> of the estimate of 5 would have to be due to bias. This implies that to
#> nullify the inference one would expect to have to replace 215 (21.506%)
#> observations with data points for which the effect is 0 (RIR = 215).
#>
#> See Frank et al. (2013) for a description of the method.
#>
#> Citation: Frank, K.A., Maroulis, S., Duong, M., and Kelcey, B. (2013).
#> What would it take to change an inference?
#> Using Rubin's causal model to interpret the robustness of causal inferences.
#> Education, Evaluation and Policy Analysis, 35 437-460.
#>
#> Accuracy of results increases with the number of decimals reported.The output provides several key metrics:
Robustness of Inference (RIR): The number of observations that would need to be replaced with observations having null effects to invalidate the inference
Percentage of sample that would need to be replaced: RIR / n_obs * 100
Impact threshold: The correlation between an omitted variable and both the treatment and outcome needed to invalidate the inference
Interpretation:
Higher RIR = more robust inference
RIR should be compared to n_obs to assess practical robustness
Impact threshold shows how strong confounding needs to be
42.4.2 Visualizing Sensitivity: The Threshold Plot
In Figure 42.9, the threshold plot shows the combination of correlations between a confound and the treatment (horizontal axis) and outcome (vertical axis) that would be needed to overturn your conclusion:
# Create threshold plot
# This visualizes the "confounding space" that would invalidate inference
pkonfound(
est_eff = 5,
std_err = 2,
n_obs = 1000,
n_covariates = 5,
to_return = "thresh_plot" # Request threshold plot
)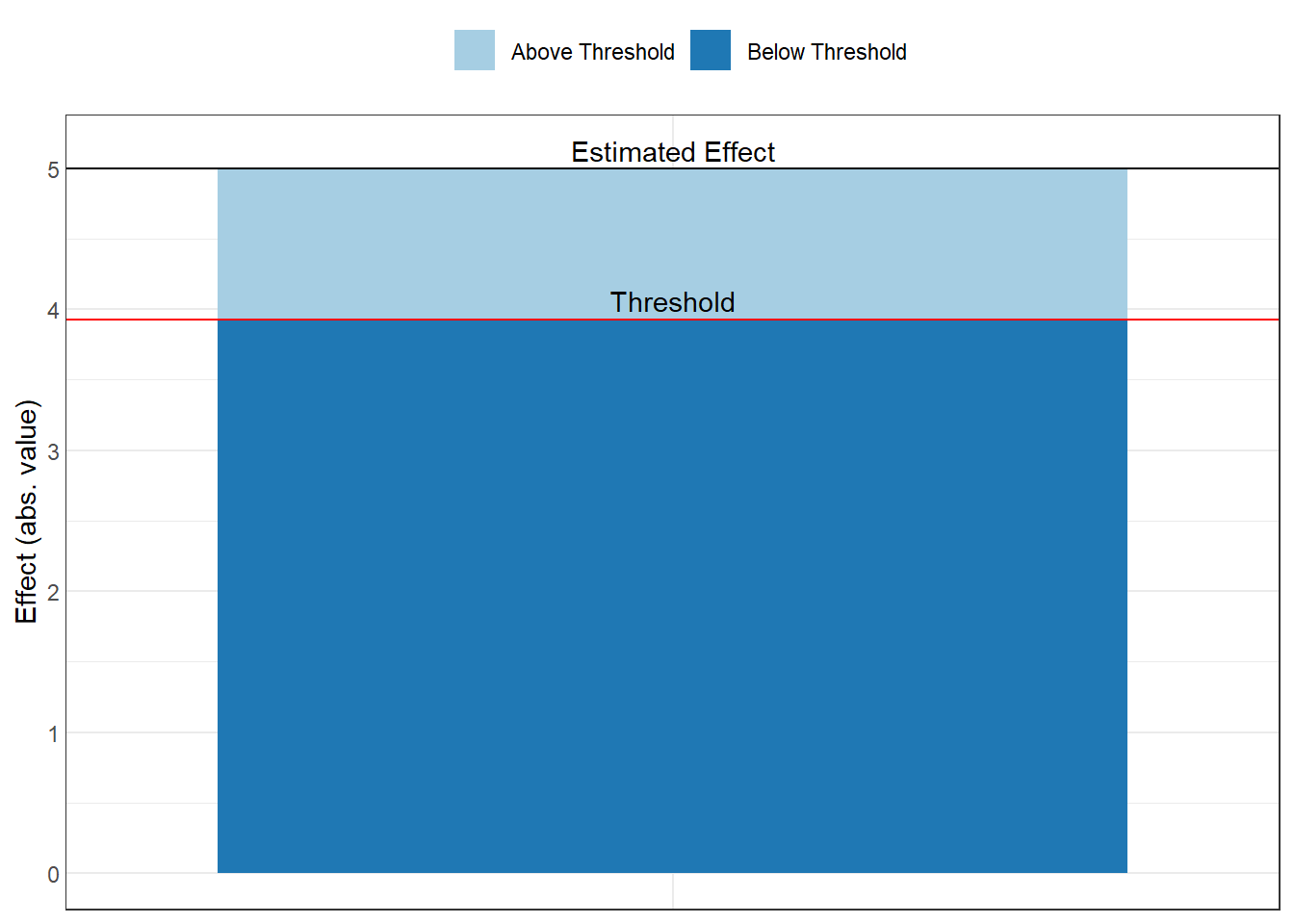
Figure 42.9: Estimated effect size relative to the robustness threshold.
Interpretation of the plot:
The red line shows the threshold
Points above/beyond this line represent confounding strong enough to overturn your inference
You can compare this to the strength of known confounds
Benchmark correlations (e.g., 0.1, 0.3, 0.5) help assess plausibility
Example interpretation: “An omitted variable would need to be correlated at 0.35 with both the treatment and outcome to invalidate our inference. Given that our strongest observed control is correlated at 0.25 with the outcome, this seems unlikely.”
42.4.3 The Correlation Plot
In Figure 42.10, the correlation plot provides another view, showing the required partial correlation of an omitted variable with the outcome, conditional on the treatment and covariates:
# Create correlation plot
pkonfound(
est_eff = 5,
std_err = 2,
n_obs = 1000,
n_covariates = 5,
to_return = "corr_plot" # Request correlation plot
)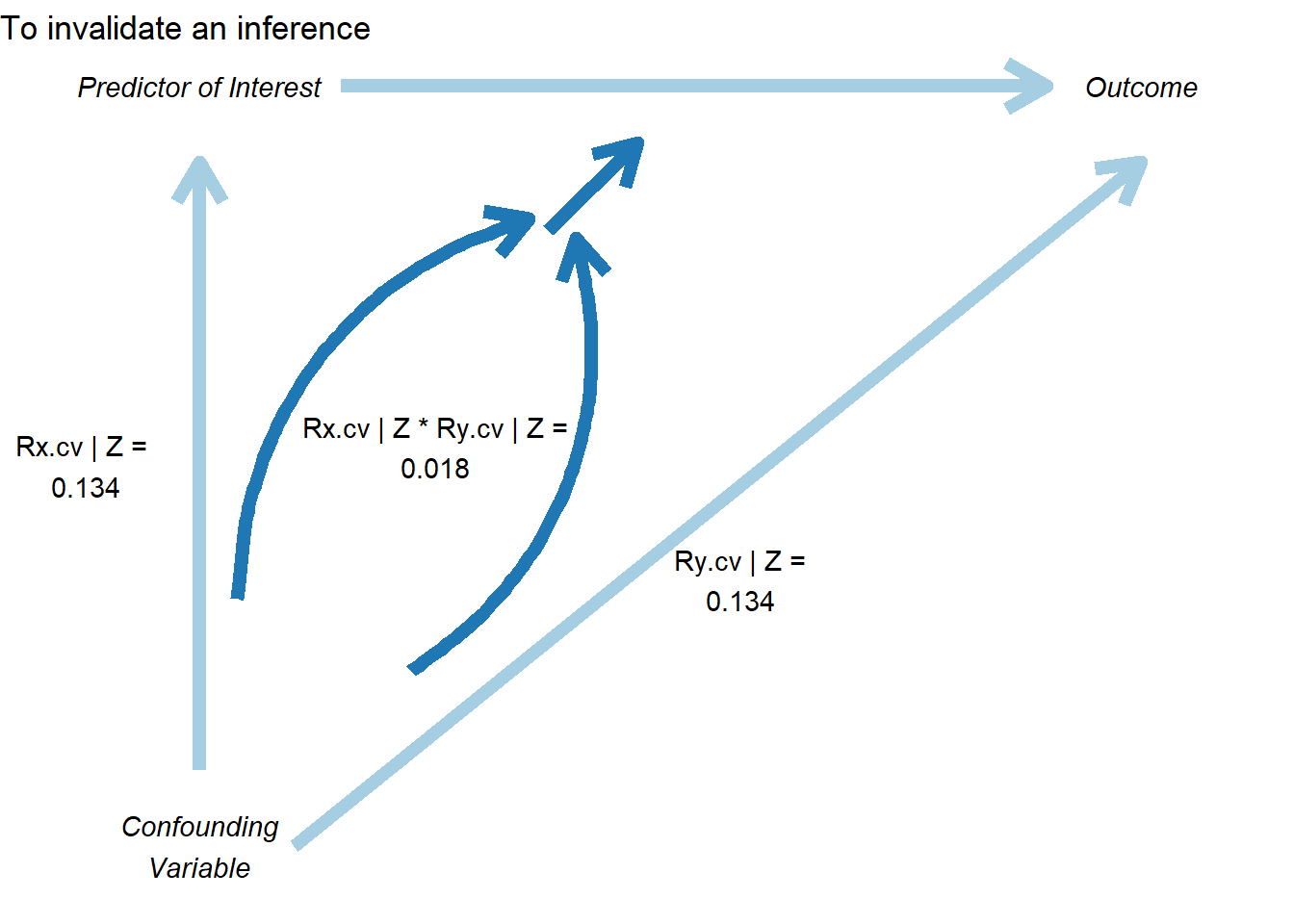
Figure 42.10: Sensitivity analysis showing conditions required to invalidate a causal inference.
This plot shows:
The relationship between bias and the required correlation
How much the effect estimate would change for different confound strengths
The threshold where the inference would be overturned
42.4.4 Konfound for Model Objects
You can also apply konfound directly to model objects, which is more convenient for real analyses:
# Fit your model
model = lm(mpg ~ wt + hp + qsec, data = mtcars)
# Apply konfound to the model
# This automatically extracts the necessary statistics
konfound(model, wt) # Assess robustness for the 'wt' coefficient
42.5 Cinelli–Hazlett Robustness Value (sensemakr)
While Oster’s \(\delta\) summarizes selection on unobservables relative to observables, and konfound rephrases the question in terms of the share of cases that would need to be replaced, Cinelli and Hazlett (2020) provide a complementary and now widely adopted answer. They ask: how strong would an unobserved confounder need to be, in terms of partial \(R^2\) with both the treatment and the outcome, before the estimated effect would lose statistical significance, or be reduced to zero? Their sensemakr package operationalizes this question with two interpretable summaries that can be reported alongside any regression-based effect estimate.
42.5.1 Motivation and the Two Key Statistics
The framework is built around partial \(R^2\) values rather than abstract selection ratios. For a hypothesized confounder \(Z\), two quantities matter:
- \(R^2_{Y \sim Z \mid D, \mathbf{X}}\), the partial \(R^2\) of the confounder with the outcome, given treatment and observed covariates, and
- \(R^2_{D \sim Z \mid \mathbf{X}}\), the partial \(R^2\) of the confounder with the treatment, given observed covariates.
A confounder is dangerous only if it is reasonably well associated with both. From these two ingredients, Cinelli and Hazlett (2020) derive two headline summaries:
- Robustness Value (RV). The smallest value \(\mathrm{RV}_q\) such that a confounder with partial \(R^2\) at least \(\mathrm{RV}_q\) on both the treatment and the outcome would be sufficient to reduce the estimated coefficient by a fraction \(q\). With \(q = 1\) this is the strength needed to drive the point estimate to zero; with \(q = q^\star\) chosen so that the bound coincides with the conventional \(t\)-statistic threshold (typically \(1.96\)), the RV is the strength needed to make the estimate statistically insignificant.
- Partial \(R^2\) of treatment with outcome. This is the worst-case bound: even a confounder that explains all residual variation in the treatment can only bias the estimate by an amount governed by \(R^2_{Y \sim D \mid \mathbf{X}}\). If this number is small, no plausible confounder, however strong, can flip the sign.
A useful rule of thumb is that an RV above \(0.1\)–\(0.2\) already represents a fairly demanding bar: an unobserved confounder would need to explain at least 10–20% of the residual variation in both the treatment and the outcome, after netting out everything in the model.
42.5.2 The Extreme-Scenario Interpretation
A common objection to sensitivity analysis is that the bounds feel abstract: what does \(R^2 = 0.15\) on the treatment actually mean? sensemakr provides two natural answers.
- Benchmarking against an observed covariate. The analyst picks a covariate (or set of covariates) already in the model and asks what would happen if the unobserved confounder were \(kd \times\) as strong as that covariate, where \(kd \in \{1, 2, 3\}\) is a multiplier. This converts an abstract partial \(R^2\) into the concrete statement “a confounder \(3\times\) as predictive as gender”.
- Extreme-confounding scenario. The procedure also reports what the estimate becomes under a worst-case confounder that explains all residual variation in the outcome, varying only its association with treatment. If the estimate survives this stress test, the result is essentially impervious to omitted variable bias of that variety.
42.5.3 Worked Example: The darfur Study
The canonical illustration in Cinelli and Hazlett (2020) uses the darfur data shipped with sensemakr, drawn from a study of attitudes toward peace among survivors of violence in Darfur. The treatment directlyharmed indicates whether the respondent was directly attacked, and the outcome peacefactor is a measure of pro-peace attitudes.
library(sensemakr)
data(darfur)
# Fit the regression
darfur_model <- lm(
peacefactor ~ directlyharmed + village + female + age +
farmer_dar + herder_dar + pastvoted + hhsize_darfur,
data = darfur
)
# Run the sensitivity analysis, benchmarking against `female`
darfur_sens <- sensemakr(
model = darfur_model,
treatment = "directlyharmed",
benchmark_covariates = "female",
kd = 1:3
)
summary(darfur_sens)
#> Sensitivity Analysis to Unobserved Confounding
#>
#> Model Formula: peacefactor ~ directlyharmed + village + female + age + farmer_dar +
#> herder_dar + pastvoted + hhsize_darfur
#>
#> Null hypothesis: q = 1 and reduce = TRUE
#> -- This means we are considering biases that reduce the absolute value of the current estimate.
#> -- The null hypothesis deemed problematic is H0:tau = 0
#>
#> Unadjusted Estimates of 'directlyharmed':
#> Coef. estimate: 0.0973
#> Standard Error: 0.0233
#> t-value (H0:tau = 0): 4.1844
#>
#> Sensitivity Statistics:
#> Partial R2 of treatment with outcome: 0.0219
#> Robustness Value, q = 1: 0.1388
#> Robustness Value, q = 1, alpha = 0.05: 0.0763
#>
#> Verbal interpretation of sensitivity statistics:
#>
#> -- Partial R2 of the treatment with the outcome: an extreme confounder (orthogonal to the covariates) that explains 100% of the residual variance of the outcome, would need to explain at least 2.19% of the residual variance of the treatment to fully account for the observed estimated effect.
#>
#> -- Robustness Value, q = 1: unobserved confounders (orthogonal to the covariates) that explain more than 13.88% of the residual variance of both the treatment and the outcome are strong enough to bring the point estimate to 0 (a bias of 100% of the original estimate). Conversely, unobserved confounders that do not explain more than 13.88% of the residual variance of both the treatment and the outcome are not strong enough to bring the point estimate to 0.
#>
#> -- Robustness Value, q = 1, alpha = 0.05: unobserved confounders (orthogonal to the covariates) that explain more than 7.63% of the residual variance of both the treatment and the outcome are strong enough to bring the estimate to a range where it is no longer 'statistically different' from 0 (a bias of 100% of the original estimate), at the significance level of alpha = 0.05. Conversely, unobserved confounders that do not explain more than 7.63% of the residual variance of both the treatment and the outcome are not strong enough to bring the estimate to a range where it is no longer 'statistically different' from 0, at the significance level of alpha = 0.05.
#>
#> Bounds on omitted variable bias:
#>
#> --The table below shows the maximum strength of unobserved confounders with association with the treatment and the outcome bounded by a multiple of the observed explanatory power of the chosen benchmark covariate(s).
#>
#> Bound Label R2dz.x R2yz.dx Treatment Adjusted Estimate Adjusted Se
#> 1x female 0.0092 0.1246 directlyharmed 0.0752 0.0219
#> 2x female 0.0183 0.2493 directlyharmed 0.0529 0.0204
#> 3x female 0.0275 0.3741 directlyharmed 0.0304 0.0187
#> Adjusted T Adjusted Lower CI Adjusted Upper CI
#> 3.4389 0.0323 0.1182
#> 2.6002 0.0130 0.0929
#> 1.6281 -0.0063 0.0670The summary output reports the unadjusted point estimate and standard error, the robustness value to bring the estimate to zero, the robustness value to bring the \(t\)-statistic below the conventional significance threshold, and the partial \(R^2\) of treatment with outcome. It then tabulates the bias-adjusted estimate, \(t\)-statistic, and lower confidence bound under hypothetical confounders that are \(1\times\), \(2\times\), and \(3\times\) as strong as the chosen benchmark covariate (female in this example).
For the Darfur data, the RV is on the order of \(0.14\), meaning that a confounder would need a partial \(R^2\) of about 14% with both treatment and outcome to drive the estimate to zero. That is substantially stronger than the role played by female in this regression, which lends credibility to the original conclusion.
42.5.4 The Sensitivity Contour Plot
The most informative output of sensemakr is a contour plot in the partial-\(R^2\) plane (Fig. 42.11). The horizontal axis represents the partial \(R^2\) of the confounder with the treatment; the vertical axis represents the partial \(R^2\) of the confounder with the outcome. Contour lines show the bias-adjusted estimate (or \(t\)-statistic) for every combination of the two. Points labeled \(1\times\), \(2\times\), \(3\times\) indicate where confounders of the corresponding multiples of the benchmark covariate would land in this plane. If those points sit far inside the region of “estimates that remain significant”, the result is robust.
plot(darfur_sens)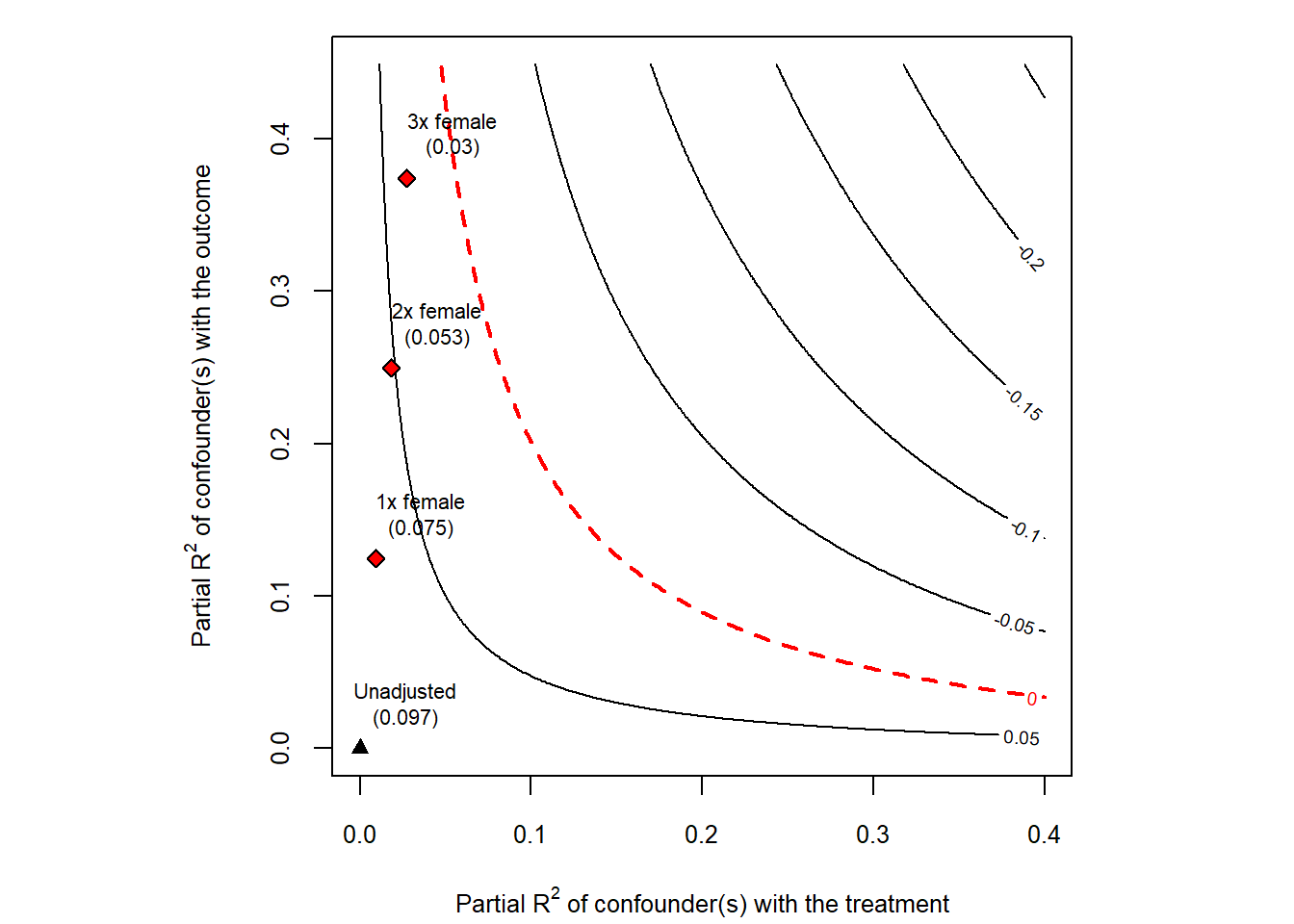
Figure 42.11: Cinelli and Hazlett sensitivity contour plot for the darfur regression.
The same machinery can be used to plot the bias-adjusted \(t\)-statistic instead of the point estimate, by passing sensitivity.of = "t-value" to plot. This is convenient when the practical question is whether a confounder of a given strength would cause the result to lose statistical significance.
42.5.5 Practical Recommendations
A few practical points are worth keeping in mind:
- Pick benchmarks that are substantively informative. Choose covariates that are strong predictors of both treatment and outcome and whose role in the system is well understood, so that the \(kd\)-multiplier statements are meaningful.
-
Report RV alongside the point estimate. A coefficient of
0.10withRV = 0.20andRV (alpha = 0.05) = 0.13tells the reader much more than a coefficient and a \(p\)-value alone. -
Combine with other tools. RV/
sensemakris complementary to Oster’s \(\delta\) (which gives a single scalar),konfound(which expresses the threshold as a share of biased cases), and the matching-based Rosenbaum bounds described next.
42.6 Rosenbaum Bounds
The methods discussed so far are designed for regression-based estimators. In matched observational studies, where the analyst has explicitly paired treated units with similar controls on observed covariates, a different and arguably even more natural sensitivity tool is available: the Rosenbaum (2002b) sensitivity bound. It asks how much the matched pairs would have to differ on an unobserved covariate before the conclusions would be overturned.
42.6.1 Motivation and the \(\Gamma\) Parameter
After matching, two units in the same matched set look identical on observed covariates \(\mathbf{X}\). If the matching were perfect and treatment assignment were ignorable given \(\mathbf{X}\), then within each matched set treatment would be assigned by a coin flip. Rosenbaum’s framework asks: what if the coin is biased, and the bias comes from an unobserved covariate \(U\) that we did not match on?
Formally, for two units \(i\) and \(j\) in the same matched set with treatment probabilities \(\pi_i\) and \(\pi_j\), the odds ratio is bounded by
\[ \frac{1}{\Gamma} \;\le\; \frac{\pi_i / (1 - \pi_i)}{\pi_j / (1 - \pi_j)} \;\le\; \Gamma. \]
The parameter \(\Gamma \ge 1\) measures the maximum departure from random assignment that an unobserved confounder could induce.
- \(\Gamma = 1\) corresponds to a randomized experiment: matched units are equally likely to be treated.
- \(\Gamma = 2\) means that, within a matched pair, one unit could be up to twice as likely as its partner to be treated, on the basis of some unobserved difference.
- \(\Gamma = 5\) would mean the unobserved confounder has enormous leverage over treatment assignment.
The sensitivity analysis proceeds by computing, at each value of \(\Gamma\), an upper bound on the \(p\)-value (or a lower bound on the test statistic) for the treatment effect under the worst-case allocation of \(U\) across pairs. The critical \(\Gamma\) is the value at which the bound first crosses the chosen significance threshold.
A widely cited rule of thumb is that critical \(\Gamma\) values above about \(2\) generally indicate a robust finding for a binary treatment in social-science contexts: an unobserved confounder would need to roughly double the odds of treatment to overturn the result. Examples of \(\Gamma\) thresholds in published studies are tabulated in Table 35.1 of the matching-methods chapter.
42.6.2 Worked Example: Lalonde Job-Training Data
The lalonde dataset packaged with MatchIt is a standard testbed for matching methods. The treatment is participation in a job-training program, and the outcome re78 is real earnings in 1978. We construct a nearest-neighbor match on a small set of pre-treatment covariates and then compute Rosenbaum bounds on the matched pairs.
library(MatchIt)
library(rbounds)
data("lalonde", package = "MatchIt")
# 1:1 nearest-neighbor matching on observed covariates
m_out <- matchit(
treat ~ age + educ + race + married + nodegree + re74 + re75,
data = lalonde,
method = "nearest"
)
matched <- match.data(m_out)
# Wilcoxon signed-rank Rosenbaum bounds
psens(
matched$re78[matched$treat == 1],
matched$re78[matched$treat == 0],
Gamma = 2,
GammaInc = 0.1
)
#>
#> Rosenbaum Sensitivity Test for Wilcoxon Signed Rank P-Value
#>
#> Unconfounded estimate .... 0.1928
#>
#> Gamma Lower bound Upper bound
#> 1.0 0.1928 0.1928
#> 1.1 0.0796 0.3709
#> 1.2 0.0285 0.5640
#> 1.3 0.0091 0.7300
#> 1.4 0.0026 0.8489
#> 1.5 0.0007 0.9226
#> 1.6 0.0002 0.9633
#> 1.7 0.0000 0.9837
#> 1.8 0.0000 0.9932
#> 1.9 0.0000 0.9973
#> 2.0 0.0000 0.9990
#>
#> Note: Gamma is Odds of Differential Assignment To
#> Treatment Due to Unobserved Factors
#> The psens() function returns, for each value of \(\Gamma\) between \(1\) and the supplied upper limit, a lower and upper \(p\)-value bound for the Wilcoxon signed-rank test under the worst-case unobserved confounder. The critical \(\Gamma\) is the smallest value at which the upper \(p\)-value bound exceeds the chosen significance level (often \(0.05\)).
For Hodges–Lehmann point and interval estimates of the treatment effect under the same family of bounds, rbounds::hlsens() provides the analogous sensitivity table:
hlsens(
matched$re78[matched$treat == 1],
matched$re78[matched$treat == 0],
Gamma = 2,
GammaInc = 0.1
)
#>
#> Rosenbaum Sensitivity Test for Hodges-Lehmann Point Estimate
#>
#> Unconfounded estimate .... 686.1615
#>
#> Gamma Lower bound Upper bound
#> 1.0 686.160 686.16
#> 1.1 165.060 932.06
#> 1.2 -50.038 1308.00
#> 1.3 -407.340 1581.90
#> 1.4 -680.640 1853.70
#> 1.5 -938.240 2039.30
#> 1.6 -1219.100 2268.00
#> 1.7 -1452.600 2514.10
#> 1.8 -1716.700 2756.70
#> 1.9 -1960.600 3070.00
#> 2.0 -2185.800 3282.40
#>
#> Note: Gamma is Odds of Differential Assignment To
#> Treatment Due to Unobserved Factors
#> 42.6.3 Interpreting and Reporting
When reporting Rosenbaum bounds, three numbers are typically of interest:
- the unadjusted test statistic and \(p\)-value (corresponding to \(\Gamma = 1\)),
- the critical \(\Gamma\) at which the upper \(p\)-value bound reaches the chosen significance threshold, and
- the bias-adjusted point and interval estimates from
hlsens()at one or two interpretable values of \(\Gamma\) (for example \(\Gamma = 1.5\) and \(\Gamma = 2\)).
It is good practice to interpret the critical \(\Gamma\) in domain-specific language: “to overturn this conclusion, an unobserved confounder would need to make program participants up to \(\Gamma\) times more likely to be matched into the treatment group than their observably similar controls, even after conditioning on age, education, race, marital status, schooling, and prior earnings.” Whether such a confounder is plausible is a substantive judgment that the bound itself cannot settle.
42.6.4 Extensions
The basic Rosenbaum bound has been extended in several directions that are increasingly relevant in modern observational work:
- Clustered Rosenbaum bounds (B. B. Hansen, Rosenbaum, and Small 2014) adapt the framework to settings where treatment is assigned at a higher level than the unit of analysis (for example, treatment at the school level with student outcomes), so that biases need to be evaluated at the cluster level rather than pair by pair.
- Amplification and the \((\Lambda, \Delta)\) parameterization, due to Rosenbaum and Silber (2009), decomposes the single sensitivity parameter \(\Gamma\) into two more interpretable pieces: \(\Lambda\), the maximum impact of \(U\) on treatment, and \(\Delta\), the maximum impact of \(U\) on the outcome. A given \(\Gamma\) corresponds to a one-dimensional curve in the \((\Lambda, \Delta)\) plane, which can be plotted alongside the Cinelli–Hazlett contour from the previous section.
-
Sensitivity for matched comparative studies with multiple control groups, instruments, or regression discontinuity-style designs is implemented in
sensitivitymv,sensitivitymw, and related packages, which generalizerboundsto weighted M-estimators and to studies with more than one matched control per treated unit.
In practice, the Rosenbaum framework and the Cinelli–Hazlett RV framework are usually run side by side. They probe overlapping but distinct aspects of unmeasured confounding: \(\Gamma\) speaks the language of treatment-assignment odds within matched sets, while RV speaks the language of partial \(R^2\) in the regression that produced the point estimate. Reporting both gives the reader a much more complete picture of how fragile or robust a causal claim really is.
42.7 Advanced Topics in Sensitivity Analysis
42.7.1 Sensitivity to Outliers and Influential Observations
Outliers can drive results, so it’s important to assess robustness to their inclusion:
# Identify influential observations
model = lm(mpg ~ wt + hp + qsec, data = mtcars)
# Calculate influence measures
influence_measures = influence.measures(model)
# Cook's Distance
cooks_d = cooks.distance(model)
influential = cooks_d > 4 / nrow(mtcars) # Common threshold
# DFBETAS (change in coefficient when observation removed)
dfbetas_vals = dfbetas(model)
# Compare models with/without influential observations
model_full = lm(mpg ~ wt + hp + qsec, data = mtcars)
model_no_influential = lm(
mpg ~ wt + hp + qsec,
data = mtcars[!influential, ]
)
# Compare coefficients
compare_df = data.frame(
Variable = names(coef(model_full)),
Full_Sample = coef(model_full),
Excl_Influential = c(
coef(model_no_influential),
rep(NA, length(coef(model_full)) - length(coef(model_no_influential)))
)
)
# Winsorization approach
library(DescTools)
mtcars_winsor = mtcars
mtcars_winsor$wt = DescTools::Winsorize(mtcars$wt, val = c(0.01, 0.99))
mtcars_winsor$hp = DescTools::Winsorize(mtcars$hp, val = c(0.01, 0.99))
model_winsor = lm(mpg ~ wt + hp + qsec, data = mtcars_winsor)
# Compare original vs. winsorized42.7.2 Sensitivity to Measurement Error
If key variables are measured with error, results may be biased:
# Simulate measurement error
# This helps understand potential attenuation bias
library(simex)
# SIMEX (Simulation-Extrapolation) for measurement error correction
# Assumes classical measurement error in covariates
# Fit model with x = TRUE
naive_model <- lm(mpg ~ wt + hp, data = mtcars, x = TRUE)
# Apply SIMEX
# lambda is the factor by which measurement error variance increases
simex_model <- simex(
naive_model,
SIMEXvariable = "wt", # Variable with measurement error
measurement.error = 0.1, # Assumed ME variance (proportion of var)
lambda = seq(0.1, 2, 0.1), # Extrapolation sequence
B = 100 # Number of simulations
)
# View results
summary(simex_model)
#> Call:
#> simex(model = naive_model, SIMEXvariable = "wt", measurement.error = 0.1,
#> lambda = seq(0.1, 2, 0.1), B = 100)
#>
#> Naive model:
#> lm(formula = mpg ~ wt + hp, data = mtcars, x = TRUE)
#>
#> Simex variable :
#> wt
#> Measurement error : 0.1
#>
#>
#> Number of iterations: 100
#>
#> Residuals:
#> Min. 1st Qu. Median Mean 3rd Qu. Max.
#> -3.931701 -1.603200 -0.225173 -0.007991 1.066691 5.806683
#>
#> Coefficients:
#>
#> Asymptotic variance:
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) 37.376807 1.987465 18.806 < 2e-16 ***
#> wt -3.974826 0.644667 -6.166 1.01e-06 ***
#> hp -0.030611 0.006577 -4.654 6.63e-05 ***
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
#>
#> Jackknife variance:
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) 37.376807 1.583219 23.608 < 2e-16 ***
#> wt -3.974826 0.636895 -6.241 8.24e-07 ***
#> hp -0.030611 0.009077 -3.372 0.00213 **
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
# Compare naive vs corrected coefficients
data.frame(
Coefficient = names(coef(naive_model)),
Naive = coef(naive_model),
SIMEX_Corrected = coef(simex_model)
)
#> Coefficient Naive SIMEX_Corrected
#> (Intercept) (Intercept) 37.22727012 37.37680699
#> wt wt -3.87783074 -3.97482594
#> hp hp -0.03177295 -0.03061053
# SIMEX extrapolates back to zero ME, giving corrected estimateFigure 42.12 plots SIMEX extrapolation.
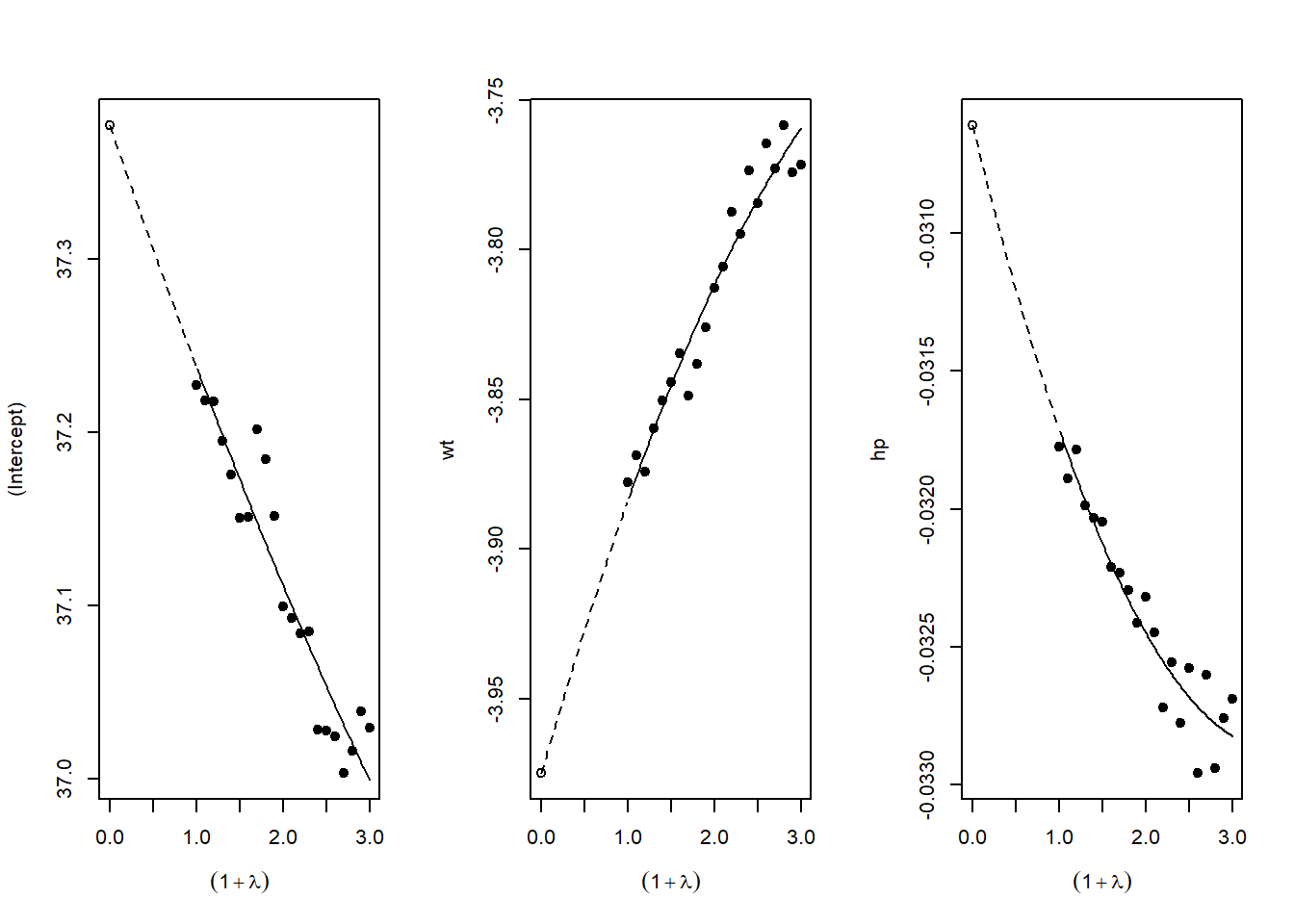
Figure 42.12: SIMEX extrapolation plots for measurement-error correction.
42.8 Reporting Sensitivity Analysis Results
42.8.1 Best Practices for Presentation
When presenting sensitivity analyses in your paper:
Main Text: Present your primary specification and 1-2 key robustness checks that directly address the most plausible threats to identification
-
Tables: Create comprehensive tables showing:
- Coefficient estimates across specifications
- Standard errors (in parentheses)
- \(R^2\) and other fit statistics
- Number of observations
- Clear column headers describing each specification
Figures: Use specification curves for visual impact when you have many specifications
Appendix: Place exhaustive robustness checks in appendices with clear organization
42.9 Conclusion and Recommendations
Sensitivity analysis is not optional in modern empirical research, it’s essential. The techniques described in this chapter provide a toolkit for assessing the robustness of your findings:
Start with specification curve analysis (
starbility) to visualize how your results vary across defensible specificationsApply Oster’s method (
robomit) to assess sensitivity to omitted variable bias using coefficient stability and \(R^2\) movementUse konfound analysis (
konfound) to quantify how much bias would be needed to overturn your inferenceConduct additional tests relevant to your context: placebo tests, subsample analysis, outlier diagnostics, etc.
Present results clearly: Use both tables and figures, provide verbal interpretation, and be transparent about which specifications you view as most credible and why
Remember: The goal is not to show that your result is robust to everything, but to demonstrate that it’s robust to reasonable alternative choices and plausible threats to identification. Be honest about the limitations while making the strongest case possible for your findings.
The mark of rigorous empirical work is not that every robustness check confirms your main result, it’s that you’ve thoughtfully considered the most important threats to validity and provided evidence about whether those threats are likely to overturn your conclusions.